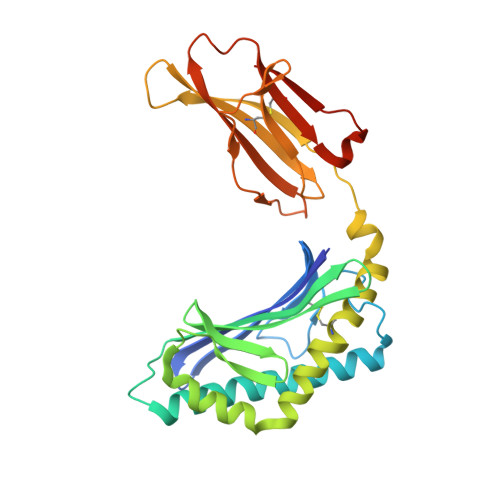
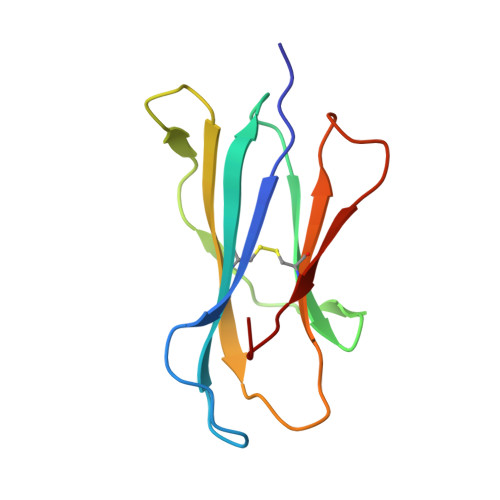
Structural evaluation of potent NKT cell agonists: implications for design of novel stimulatory ligands.
Schiefner, A., Fujio, M., Wu, D., Wong, C.H., Wilson, I.A.(2009) J Mol Biology 394: 71-82
- PubMed: 19732779 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.08.061
- Primary Citation Related Structures:
3GML, 3GMM, 3GMN, 3GMO, 3GMP, 3GMQ, 3GMR - PubMed Abstract:
Natural killer T (NKT) cells are a subset of T cells that are activated by CD1d-glycolipid complexes through a semi-invariant alphabeta T cell receptor (NKT TCR). Upon activation, NKT cells secrete regulatory cytokines that are implicated in T helper cell responses. alpha-Galactosylceramide (alpha-GalCer) is a potent NKT cell agonist when presented by CD1d. Phenyl ring substitutions of the alpha-GalCer fatty acid moiety were recently found to be superior in eliciting regulatory cytokines. Crystal structures of four new mouse CD1d-lipid complexes (five structures), a new PBS-25 complex, and CD1d with an endogenous ligand, at 1.6-1.9 A resolution, reveal that the alpha-GalCer phenyl analogues impart minor structural differences to the A'-pocket, while the sphingosine and galactose moieties, important for NKT TCR recognition, remain virtually unchanged. The observed differences in cytokine-release profiles appear to be associated with increased stability of the CD1d-glycolipid complexes rather than increased affinity for the NKT TCR. Furthermore, comparison of mouse CD1d-glycolipid complexes in different crystallographic space groups reveals considerable conformational variation, particularly above the F'-pocket, the primary site of interaction with the NKT TCR. We propose that modifications of the sphingosine moiety or other substitutions that decrease alpha-GalCer flexibility would stabilize the F'-pocket. Such compounds might then increase CD1d affinity for the NKT TCR and further enhance the stimulatory and regulatory properties of alpha-GalCer derivatives.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: