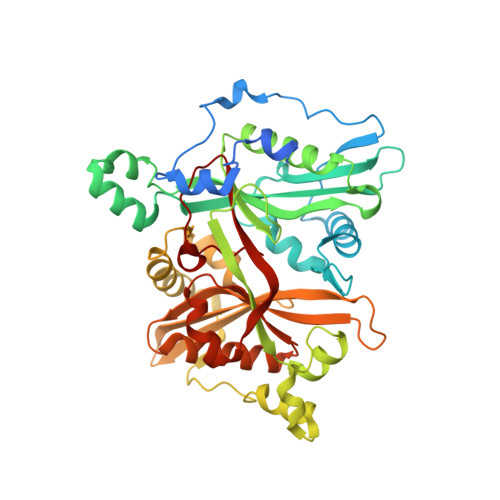
N-myristoyltransferase inhibitors as new leads to treat sleeping sickness.
Frearson, J.A., Brand, S., McElroy, S.P., Cleghorn, L.A., Smid, O., Stojanovski, L., Price, H.P., Guther, M.L., Torrie, L.S., Robinson, D.A., Hallyburton, I., Mpamhanga, C.P., Brannigan, J.A., Wilkinson, A.J., Hodgkinson, M., Hui, R., Qiu, W., Raimi, O.G., van Aalten, D.M., Brenk, R., Gilbert, I.H., Read, K.D., Fairlamb, A.H., Ferguson, M.A., Smith, D.F., Wyatt, P.G.(2010) Nature 464: 728-732
- PubMed: 20360736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature08893
- Primary Citation Related Structures:
2WSA, 3H5Z - PubMed Abstract:
African sleeping sickness or human African trypanosomiasis, caused by Trypanosoma brucei spp., is responsible for approximately 30,000 deaths each year. Available treatments for this disease are poor, with unacceptable efficacy and safety profiles, particularly in the late stage of the disease when the parasite has infected the central nervous system. Here we report the validation of a molecular target and the discovery of associated lead compounds with the potential to address this lack of suitable treatments. Inhibition of this target-T. brucei N-myristoyltransferase-leads to rapid killing of trypanosomes both in vitro and in vivo and cures trypanosomiasis in mice. These high-affinity inhibitors bind into the peptide substrate pocket of the enzyme and inhibit protein N-myristoylation in trypanosomes. The compounds identified have promising pharmaceutical properties and represent an opportunity to develop oral drugs to treat this devastating disease. Our studies validate T. brucei N-myristoyltransferase as a promising therapeutic target for human African trypanosomiasis.
- Drug Discovery Unit, Division of Biological Chemistry and Drug Discovery, University of Dundee, Dundee DD1 5EH, UK.
Organizational Affiliation: