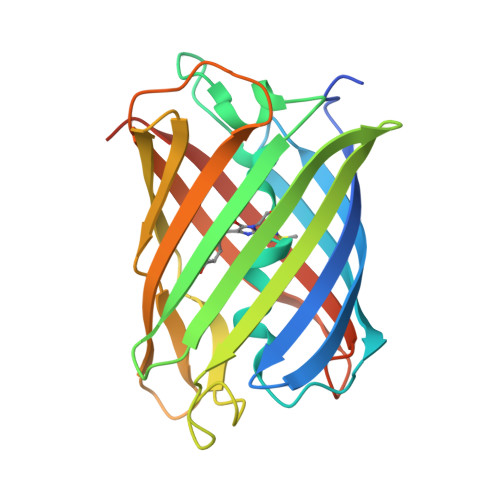
Photoactivation mechanism of PAmCherry based on crystal structures of the protein in the dark and fluorescent states.
Subach, F.V., Malashkevich, V.N., Zencheck, W.D., Xiao, H., Filonov, G.S., Almo, S.C., Verkhusha, V.V.(2009) Proc Natl Acad Sci U S A 106: 21097-21102
- PubMed: 19934036 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0909204106
- Primary Citation Related Structures:
3KCS, 3KCT - PubMed Abstract:
Photoactivatable fluorescent proteins (PAFPs) are required for super-resolution imaging of live cells. Recently, the first red PAFP, PAmCherry1, was reported, which complements the photo-activatable GFP by providing a red super-resolution color. PAmCherry1 is originally "dark" but exhibits red fluorescence after UV-violet light irradiation. To define the structural basis of PAmCherry1 photoactivation, we determined its crystal structure in the dark and red fluorescent states at 1.50 A and 1.65 A, respectively. The non-coplanar structure of the chromophore in the dark PAmChery1 suggests the presence of an N-acylimine functionality and a single non-oxidized C(alpha)-C(beta) bond in the Tyr-67 side chain in the cyclized Met-66-Tyr-67-Gly-68 tripeptide. MS data of the chromophore-bearing peptide indicates the loss of 20 Da upon maturation, whereas tandem MS reveals the C(alpha)-N bond in Met-66 is oxidized. These data indicate that PAmCherry1 in the dark state possesses the chromophore N-[(E)-(5-hydroxy-1H-imidazol-2-yl)methylidene]acetamide, which, to our knowledge, has not been previously observed in PAFPs. The photoactivated PAmCherry1 exhibits a non-coplanar anionic DsRed-like chromophore but in the trans configuration. Based on the crystallographic analysis, MS data, and biochemical analysis of the PAmCherry1 mutants, we propose the detailed photoactivation mechanism. In this mechanism, the excited-state PAmCherry1 chromophore acts as the oxidant to release CO(2) molecule from Glu-215 via a Koble-like radical reaction. The Glu-215 decarboxylation directs the carbanion formation resulting in the oxidation of the Tyr-67 C(alpha)-C(beta) bond. The double bond extends the pi-conjugation between the phenolic ring of Tyr-67, the imidazolone, and the N-acylimine, resulting in the red fluorescent chromophore.
- Department of Anatomy and Structural Biology and Gruss-Lipper Biophotonics Center, Albert Einstein College of Medicine, Bronx, NY 10461, USA.
Organizational Affiliation: