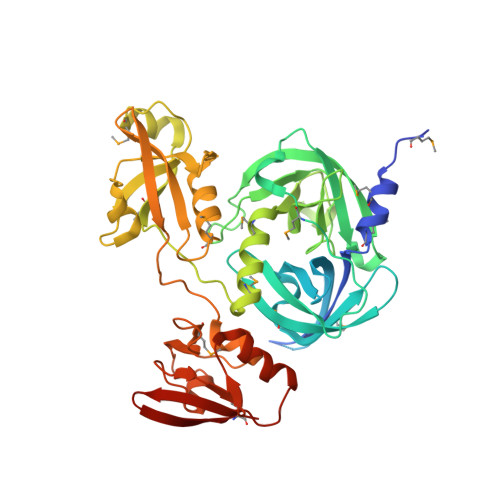
HtrA proteases have a conserved activation mechanism that can be triggered by distinct molecular cues
Krojer, T., Sawa, J., Huber, R., Clausen, T.(2010) Nat Struct Mol Biol 17: 844-852
- PubMed: 20581825 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1840
- Primary Citation Related Structures:
3MH4, 3MH5, 3MH6, 3MH7 - PubMed Abstract:
HtrA proteases are tightly regulated proteolytic assemblies that are essential for maintaining protein homeostasis in extracytosolic compartments. Though HtrA proteases have been characterized in detail, their precise molecular mechanism for switching between different functional states is still unknown. To address this, we carried out biochemical and structural studies of DegP from Escherichia coli. We show that effector-peptide binding to the PDZ domain of DegP induces oligomer conversion from resting hexameric DegP6 into proteolytically active 12-mers and 24-mers (DegP12/24). Moreover, our data demonstrate that a specific protease loop (L3) functions as a conserved molecular switch of HtrA proteases. L3 senses the activation signal-that is, the repositioned PDZ domain of substrate-engaged DegP12/24 or the binding of allosteric effectors to regulatory HtrA proteases such as DegS-and transmits this information to the active site. Implications for protein quality control and regulation of oligomeric enzymes are discussed.
- Research Institute of Molecular Pathology, Vienna, Austria.
Organizational Affiliation: