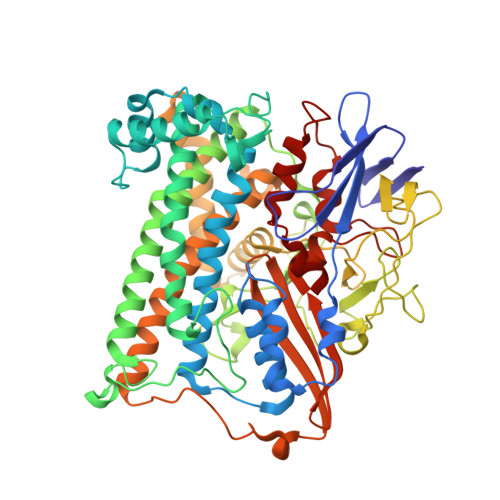
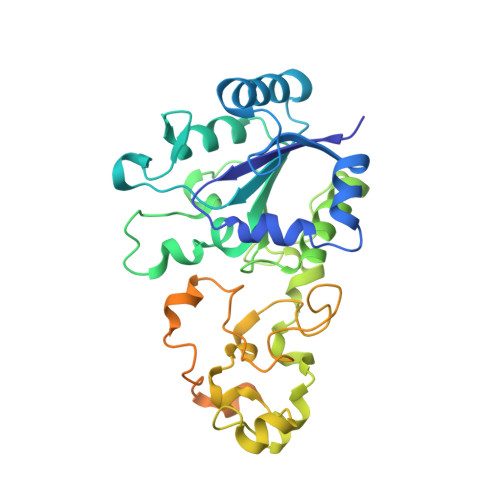
The crystal structure of an oxygen-tolerant hydrogenase uncovers a novel iron-sulphur centre.
Fritsch, J., Scheerer, P., Frielingsdorf, S., Kroschinsky, S., Friedrich, B., Lenz, O., Spahn, C.M.(2011) Nature 479: 249-252
- PubMed: 22002606 Search on PubMed
- DOI: https://doi.org/10.1038/nature10505
- Primary Citation Related Structures:
3RGW - PubMed Abstract:
Hydrogenases are abundant enzymes that catalyse the reversible interconversion of H(2) into protons and electrons at high rates. Those hydrogenases maintaining their activity in the presence of O(2) are considered to be central to H(2)-based technologies, such as enzymatic fuel cells and for light-driven H(2) production. Despite comprehensive genetic, biochemical, electrochemical and spectroscopic investigations, the molecular background allowing a structural interpretation of how the catalytic centre is protected from irreversible inactivation by O(2) has remained unclear. Here we present the crystal structure of an O(2)-tolerant [NiFe]-hydrogenase from the aerobic H(2) oxidizer Ralstonia eutropha H16 at 1.5 Å resolution. The heterodimeric enzyme consists of a large subunit harbouring the catalytic centre in the H(2)-reduced state and a small subunit containing an electron relay consisting of three different iron-sulphur clusters. The cluster proximal to the active site displays an unprecedented [4Fe-3S] structure and is coordinated by six cysteines. According to the current model, this cofactor operates as an electronic switch depending on the nature of the gas molecule approaching the active site. It serves as an electron acceptor in the course of H(2) oxidation and as an electron-delivering device upon O(2) attack at the active site. This dual function is supported by the capability of the novel iron-sulphur cluster to adopt three redox states at physiological redox potentials. The second structural feature is a network of extended water cavities that may act as a channel facilitating the removal of water produced at the [NiFe] active site. These discoveries will have an impact on the design of biological and chemical H(2)-converting catalysts that are capable of cycling H(2) in air.
- Mikrobiologie, Institut für Biologie, Humboldt-Universität zu Berlin, Chausseestraße 117, 10115 Berlin, Germany.
Organizational Affiliation: