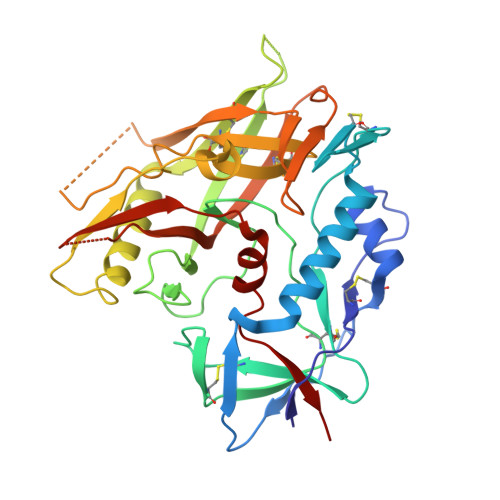
Unliganded HIV-1 gp120 core structures assume the CD4-bound conformation with regulation by quaternary interactions and variable loops.
Kwon, Y.D., Finzi, A., Wu, X., Dogo-Isonagie, C., Lee, L.K., Moore, L.R., Schmidt, S.D., Stuckey, J., Yang, Y., Zhou, T., Zhu, J., Vicic, D.A., Debnath, A.K., Shapiro, L., Bewley, C.A., Mascola, J.R., Sodroski, J.G., Kwong, P.D.(2012) Proc Natl Acad Sci U S A 109: 5663-5668
- PubMed: 22451932 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1112391109
- Primary Citation Related Structures:
3TGQ, 3TGR, 3TGS, 3TGT, 3TIH - PubMed Abstract:
The HIV-1 envelope (Env) spike (gp120(3)/gp41(3)) undergoes considerable structural rearrangements to mediate virus entry into cells and to evade the host immune response. Engagement of CD4, the primary human receptor, fixes a particular conformation and primes Env for entry. The CD4-bound state, however, is prone to spontaneous inactivation and susceptible to antibody neutralization. How does unliganded HIV-1 maintain CD4-binding capacity and regulate transitions to the CD4-bound state? To define this mechanistically, we determined crystal structures of unliganded core gp120 from HIV-1 clades B, C, and E. Notably, all of these unliganded HIV-1 structures resembled the CD4-bound state. Conformational fixation with ligand selection and thermodynamic analysis of full-length and core gp120 interactions revealed that the tendency of HIV-1 gp120 to adopt the CD4-bound conformation was restrained by the V1/V2- and V3-variable loops. In parallel, we determined the structure of core gp120 in complex with the small molecule, NBD-556, which specifically recognizes the CD4-bound conformation of gp120. Neutralization by NBD-556 indicated that Env spikes on primary isolates rarely assume the CD4-bound conformation spontaneously, although they could do so when quaternary restraints were loosened. Together, the results suggest that the CD4-bound conformation represents a "ground state" for the gp120 core, with variable loop and quaternary interactions restraining unliganded gp120 from "snapping" into this conformation. A mechanism of control involving deformations in unliganded structure from a functionally critical state (e.g., the CD4-bound state) provides advantages in terms of HIV-1 Env structural diversity and resistance to antibodies and inhibitors, while maintaining elements essential for entry.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: