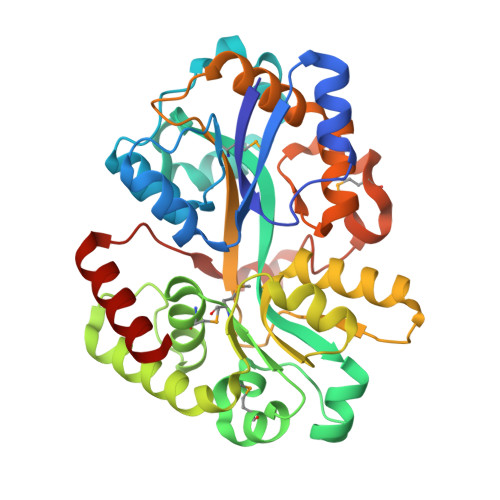
Structural Basis of Substrate Binding Specificity Revealed by the Crystal Structures of Polyamine Receptors SpuD and SpuE from Pseudomonas aeruginosa
Wu, D.H., Lim, S.C., Dong, Y.H., Wu, J.E., Tao, F., Zhou, L., Zhang, L.H., Song, H.W.(2012) J Mol Biology 416: 697-712
- PubMed: 22300763 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.01.010
- Primary Citation Related Structures:
3TTK, 3TTL, 3TTM, 3TTN - PubMed Abstract:
The type III secretion system (T3SS) of Pseudomonas aeruginosa is a key virulence determinant whose expression is induced by polyamine signals from mammalian host. SpuD and SpuE were postulated to be spermidine-preferential binding proteins, which regulate the polyamine content in this bacterial pathogen. In this study, we found that SpuD is a putrescine-preferential binding protein, while SpuE binds to spermidine exclusively. We have determined the crystal structures of SpuD in free form and in complex with putrescine and SpuE in free form and in complex with spermidine. Upon ligand binding, SpuD and SpuE undergo an "open-to-closed" conformational switch with the resultant closed ligand-bound forms, SpuD-putrescine and SpuE-spermidine, similar to their Escherichia coli counterparts PotF-putrescine and PotD-spermidine, respectively. Structural comparison suggested that two aromatic residues, Trp271 of SpuE and Phe273 of SpuD in segment II region, are the key structural determinants for putrescine/spermidine recognition specificity. Mutagenesis combined with isothermal titration calorimetry showed that substitution of Trp271 by Phe enabled SpuE to gain substantial binding affinity for putrescine, while replacement of Phe273 by Trp reduced the binding affinity of SpuD toward putrescine by 250-fold. Altogether, these results revealed the molecular mechanism governing polyamine recognition specificity by SpuD and SpuE and provide the basis for further structural and functional studies of polyamine signal importation system in P. aeruginosa.
- Institute of Molecular and Cell Biology, 61 Biopolis Drive, Proteos, Singapore 138673.
Organizational Affiliation: