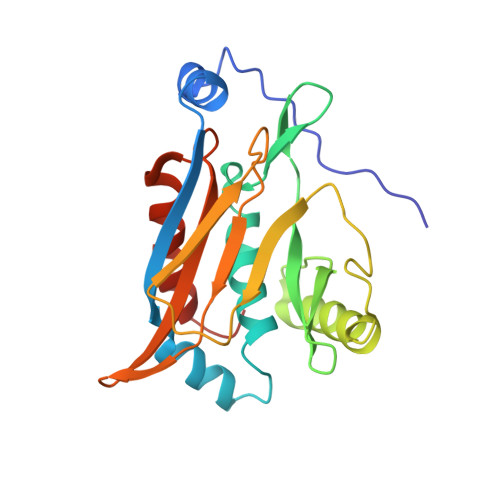
The structural basis for substrate versatility of chloramphenicol acetyltransferase CAT(I).
Biswas, T., Houghton, J.L., Garneau-Tsodikova, S., Tsodikov, O.V.(2012) Protein Sci 21: 520-530
- PubMed: 22294317 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2036
- Primary Citation Related Structures:
3U9B, 3U9F - PubMed Abstract:
Novel antibiotics are needed to overcome the challenge of continually evolving bacterial resistance. This has led to a renewed interest in mechanistic studies of once popular antibiotics like chloramphenicol (CAM). Chloramphenicol acetyltransferases (CATs) are enzymes that covalently modify CAM, rendering it inactive against its target, the ribosome, and thereby causing resistance to CAM. Of the three major types of CAT (CAT(I-III)), the CAM-specific CAT(III) has been studied extensively. Much less is known about another clinically important type, CAT(I). In addition to inactivating CAM and unlike CAT(III), CAT(I) confers resistance to a structurally distinct antibiotic, fusidic acid. The origin of the broader substrate specificity of CAT(I) has not been fully elucidated. To understand the substrate binding features of CAT(I), its crystal structures in the unbound (apo) and CAM-bound forms were determined. The analysis of these and previously determined CAT(I)-FA and CAT(III)-CAM structures revealed interactions responsible for CAT(I) binding to its substrates and clarified the broader substrate preference of CAT(I) compared to that of CAT(III).
- Department of Medicinal Chemistry, University of Michigan, Ann Arbor, Michigan 48109, USA.
Organizational Affiliation: