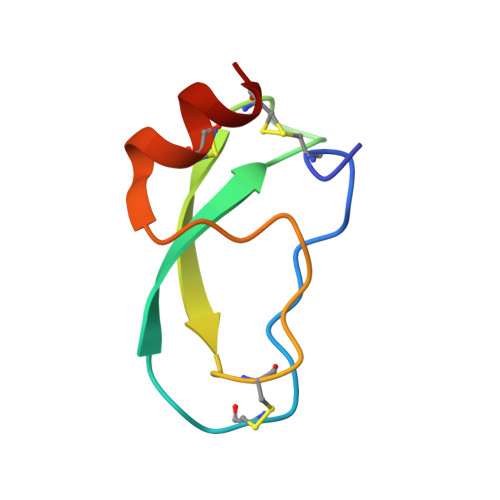
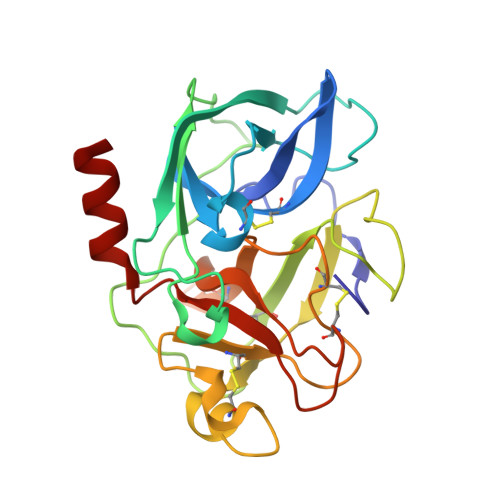
Three-dimensional Structure of a Kunitz-type Inhibitor in Complex with an Elastase-like Enzyme.
Garcia-Fernandez, R., Perbandt, M., Rehders, D., Ziegelmuller, P., Piganeau, N., Hahn, U., Betzel, C., Chavez, M.A., Redecke, L.(2015) J Biological Chem 290: 14154-14165
- PubMed: 25878249 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.647586
- Primary Citation Related Structures:
3UOU - PubMed Abstract:
Elastase-like enzymes are involved in important diseases such as acute pancreatitis, chronic inflammatory lung diseases, and cancer. Structural insights into their interaction with specific inhibitors will contribute to the development of novel anti-elastase compounds that resist rapid oxidation and proteolysis. Proteinaceous Kunitz-type inhibitors homologous to the bovine pancreatic trypsin inhibitor (BPTI) provide a suitable scaffold, but the structural aspects of their interaction with elastase-like enzymes have not been elucidated. Here, we increased the selectivity of ShPI-1, a versatile serine protease inhibitor from the sea anemone Stichodactyla helianthus with high biomedical and biotechnological potential, toward elastase-like enzymes by substitution of the P1 residue (Lys(13)) with leucine. The variant (rShPI-1/K13L) exhibits a novel anti-porcine pancreatic elastase (PPE) activity together with a significantly improved inhibition of human neuthrophil elastase and chymotrypsin. The crystal structure of the PPE·rShPI-1/K13L complex determined at 2.0 Å resolution provided the first details of the canonical interaction between a BPTI-Kunitz-type domain and elastase-like enzymes. In addition to the essential impact of the variant P1 residue for complex stability, the interface is improved by increased contributions of the primary and secondary binding loop as compared with similar trypsin and chymotrypsin complexes. A comparison of the interaction network with elastase complexes of canonical inhibitors from the chelonian in family supports a key role of the P3 site in ShPI-1 in directing its selectivity against pancreatic and neutrophil elastases. Our results provide the structural basis for site-specific mutagenesis to further improve the binding affinity and/or direct the selectivity of BPTI-Kunitz-type inhibitors toward elastase-like enzymes.
- From the Centro de Estudio de Proteínas, Facultad de Biología, Universidad de la Habana, 20146 Habana, Cuba.
Organizational Affiliation: