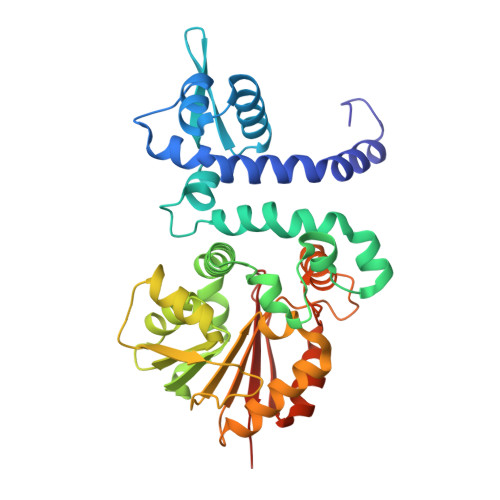
Crystal Structure and Functional Mapping of Human Asmt, the Last Enzyme of the Melatonin Synthesis Pathway.
Botros, H.G., Legrand, P., Pagan, C., Bondet, V., Weber, P., Ben-Abdallah, M., Lemiere, N., Huguet, G., Bellalou, J., Maronde, E., Beguin, P., Haouz, A., Shepard, W., Bourgeron, T.(2013) J Pineal Res 54: 46
- PubMed: 22775292 Search on PubMed
- DOI: https://doi.org/10.1111/j.1600-079X.2012.01020.x
- Primary Citation Related Structures:
4A6D, 4A6E - PubMed Abstract:
Melatonin is a synchronizer of many physiological processes. Abnormal melatonin signaling is associated with human disorders related to sleep, metabolism, and neurodevelopment. Here, we present the X-ray crystal structure of human N-acetyl serotonin methyltransferase (ASMT), the last enzyme of the melatonin biosynthesis pathway. The polypeptide chain of ASMT consists of a C-terminal domain, which is typical of other SAM-dependent O-methyltransferases, and an N-terminal domain, which intertwines several helices with another monomer to form the physiologically active dimer. Using radioenzymology, we analyzed 20 nonsynonymous variants identified through the 1000 genomes project and in patients with neuropsychiatric disorders. We found that the majority of these mutations reduced or abolished ASMT activity including one relatively frequent polymorphism in the Han Chinese population (N17K, rs17149149). Overall, we estimate that the allelic frequency of ASMT deleterious mutations ranges from 0.66% in Europe to 2.97% in Asia. Mapping of the variants on to the 3-dimensional structure clarifies why some are harmful and provides a structural basis for understanding melatonin deficiency in humans.
- Institut Pasteur, Human Genetics and Cognitive Functions Unit, Paris, France CNRS URA 2182 'Genes, synapses and cognition', Institut Pasteur, Paris, France University Paris Diderot, Sorbonne Paris Cité, Human Genetics and Cognitive Functions, Paris, France Synchrotron SOLEIL, L'Orme des Merisiers, Saint Aubin BP48, Gif-sur-Yvette, France Institut Pasteur, Plate forme 5, 25 rue Dr. Roux, Paris, France Institut Pasteur, Plate forme 6, CNRS-UMR3528, 25 rue Dr. Roux, Paris, France Institute for Anatomy III, Goethe University, Frankfurt, Germany.
Organizational Affiliation: