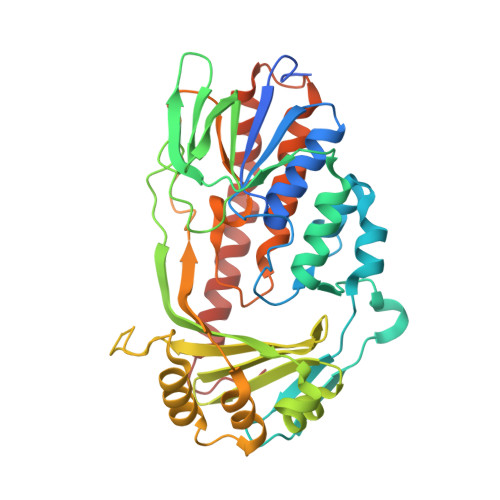
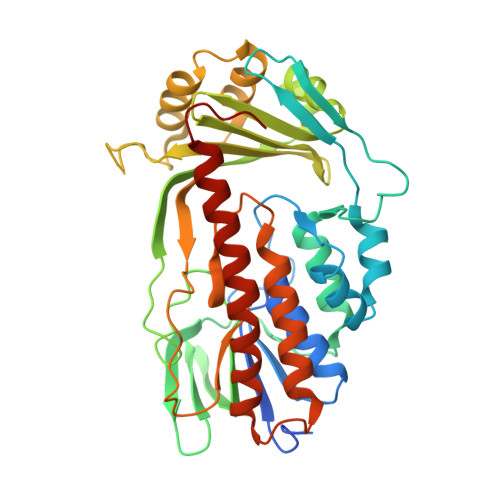
Putative Dioxygen-Binding Sites and Recognition of Tigecycline and Minocycline in the Tetracycline-Degrading Monooxygenase Tetx
Volkers, G., Damas, J.M., Palm, G.J., Panjikar, S., Soares, C.M., Hinrichs, W.(2013) Acta Crystallogr D Biol Crystallogr 69: 1758
- PubMed: 23999299 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913013802
- Primary Citation Related Structures:
4A6N, 4A99, 4GUV - PubMed Abstract:
Expression of the aromatic hydroxylase TetX under aerobic conditions confers bacterial resistance against tetracycline antibiotics. Hydroxylation inactivates and degrades tetracyclines, preventing inhibition of the prokaryotic ribosome. X-ray crystal structure analyses of TetX in complex with the second-generation and third-generation tetracyclines minocycline and tigecycline at 2.18 and 2.30 Å resolution, respectively, explain why both clinically potent antibiotics are suitable substrates. Both tetracyclines bind in a large tunnel-shaped active site in close contact to the cofactor FAD, pre-oriented for regioselective hydroxylation to 11a-hydroxytetracyclines. The characteristic bulky 9-tert-butylglycylamido substituent of tigecycline is solvent-exposed and does not interfere with TetX binding. In the TetX-minocycline complex a second binding site for a minocycline dimer is observed close to the active-site entrance. The pocket is formed by the crystal packing arrangement on the surface of two neighbouring TetX monomers. Crystal structure analysis at 2.73 Å resolution of xenon-pressurized TetX identified two adjacent Xe-binding sites. These putative dioxygen-binding cavities are located in the substrate-binding domain next to the active site. Molecular-dynamics simulations were performed in order to characterize dioxygen-diffusion pathways to FADH2 at the active site.
- Department of Molecular Structural Biology, Institute for Biochemistry, University of Greifswald, Felix-Hausdorff-Strasse 4, Greifswald, Germany.
Organizational Affiliation: