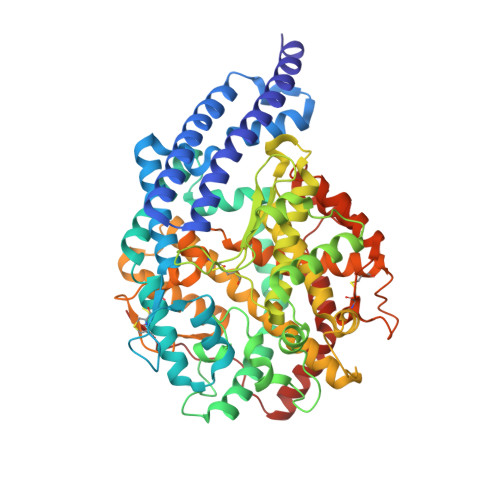
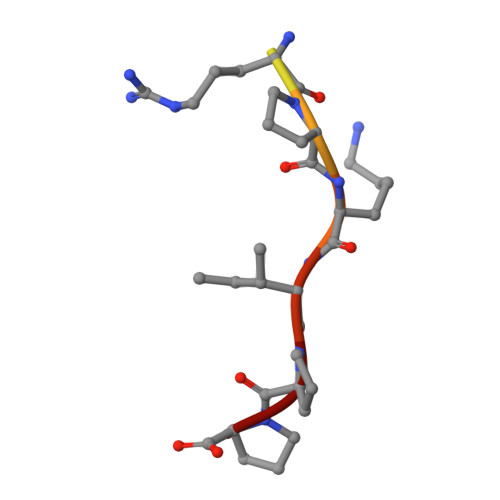
Structural Basis of Peptide Recognition by the Angiotensin-I Converting Enzyme Homologue Ance from Drosophila Melanogaster
Akif, M., Masuyer, G., Bingham, R.J., Sturrock, E.D., Isaac, R.E., Acharya, K.R.(2012) FEBS J 279: 4525
- PubMed: 23082758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.12038
- Primary Citation Related Structures:
4AA1, 4AA2, 4ASQ, 4ASR - PubMed Abstract:
Human somatic angiotensin-1 converting enzyme (ACE) is a zinc-dependent exopeptidase, that catalyses the conversion of the decapeptide angiotensin I to the octapeptide angiotensin II, by removing a C-terminal dipeptide. It is the principal component of the renin-angiotensin-aldosterone system that regulates blood pressure. Hence it is an important therapeutic target for the treatment of hypertension and cardiovascular disorders. Here, we report the structures of an ACE homologue from Drosophila melanogaster (AnCE; a proven structural model for the more complex human ACE) co-crystallized with mammalian peptide substrates (bradykinin, Thr(6) -bradykinin, angiotensin I and a snake venom peptide inhibitor, bradykinin-potentiating peptide-b). The structures determined at 2-Å resolution illustrate that both angiotensin II (the cleaved product of angiotensin I by AnCE) and bradykinin-potentiating peptide-b bind in an analogous fashion at the active site of AnCE, but also exhibit significant differences. In addition, the binding of Arg-Pro-Pro, the cleavage product of bradykinin and Thr(6) - bradykinin, provides additional detail of the general peptide binding in AnCE. Thus the new structures of AnCE complexes presented here improves our understanding of the binding of peptides and the mechanism by which peptides inhibit this family of enzymes. The atomic coordinates and structure factors for AnCE-Ang II (code 4AA1), AnCE-BPPb (code 4AA2), AnCE-BK (code 4ASQ) and AnCE-Thr6-BK (code 4ASR) complexes have been deposited in the Protein Data Bank, Research Collaboratory for Structural Bioinformatics, Rutgers University, New Brunswick, NJ (http://www.rcsb.org/) • AnCE cleaves Ang I by enzymatic study (View interaction) • Bradykinin and AnCE bind by x-ray crystallography (View interaction) • BPP and AnCE bind by x-ray crystallography (View interaction) • AnCE cleaves Bradykinin by enzymatic study (View interaction) • Ang II and AnCE bind by x-ray crystallography (View interaction).
- Department of Biology and Biochemistry, University of Bath, UK.
Organizational Affiliation: