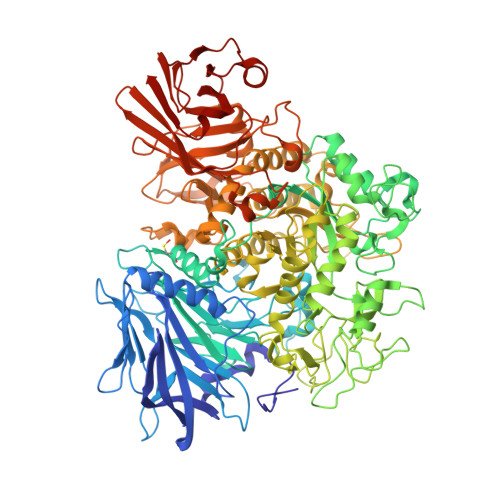
Crystal Structure of Alpha-1,4-Glucan Lyase, a Unique Glycoside Hydrolase Family Member with a Novel Catalytic Mechanism.
Rozeboom, H.J., Yu, S., Madrid, S., Kalk, K.H., Zhang, R., Dijkstra, B.W.(2013) J Biological Chem 288: 26764
- PubMed: 23902768 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.485896
- Primary Citation Related Structures:
2X2H, 2X2I, 2X2J, 4AMW, 4AMX - PubMed Abstract:
α-1,4-Glucan lyase (EC 4.2.2.13) from the red seaweed Gracilariopsis lemaneiformis cleaves α-1,4-glucosidic linkages in glycogen, starch, and malto-oligosaccharides, yielding the keto-monosaccharide 1,5-anhydro-D-fructose. The enzyme belongs to glycoside hydrolase family 31 (GH31) but degrades starch via an elimination reaction instead of hydrolysis. The crystal structure shows that the enzyme, like GH31 hydrolases, contains a (β/α)8-barrel catalytic domain with B and B' subdomains, an N-terminal domain N, and the C-terminal domains C and D. The N-terminal domain N of the lyase was found to bind a trisaccharide. Complexes of the enzyme with acarbose and 1-dexoynojirimycin and two different covalent glycosyl-enzyme intermediates obtained with fluorinated sugar analogues show that, like GH31 hydrolases, the aspartic acid residues Asp(553) and Asp(665) are the catalytic nucleophile and acid, respectively. However, as a unique feature, the catalytic nucleophile is in a position to act also as a base that abstracts a proton from the C2 carbon atom of the covalently bound subsite -1 glucosyl residue, thus explaining the unique lyase activity of the enzyme. One Glu to Val mutation in the active site of the homologous α-glucosidase from Sulfolobus solfataricus resulted in a shift from hydrolytic to lyase activity, demonstrating that a subtle amino acid difference can promote lyase activity in a GH31 hydrolase.
- From the Laboratory of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Nijenborgh 7, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: