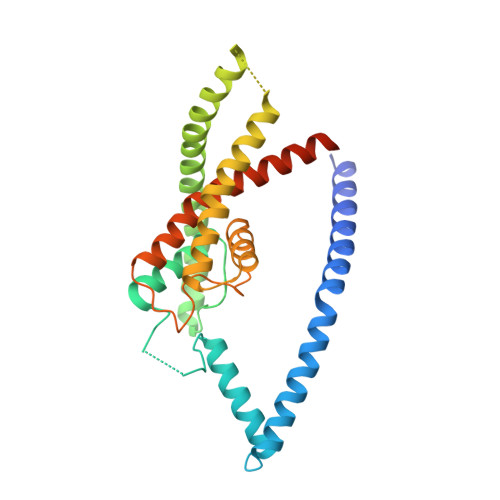
K2P Channel Gating Mechanisms Revealed by Structures of Trek-2 and a Complex with Prozac
Pike, A.C.W., Mackenzie, A., Mcclenaghan, C., Aryal, P., Dong, L., Quigley, A., Grieben, M., Goubin, S., Mukhopadhyay, S., Ruda, G.F., Clausen, M.V., Cao, L., Brennan, P.E., Burgess-Brown, N.A., Sansom, M.S.P., Tucker, S.J., Carpenter, E.P.(2015) Science 347: 1256
- PubMed: 25766236 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1261512
- Primary Citation Related Structures:
4BW5, 4XDJ, 4XDK, 4XDL - PubMed Abstract:
TREK-2 (KCNK10/K2P10), a two-pore domain potassium (K2P) channel, is gated by multiple stimuli such as stretch, fatty acids, and pH and by several drugs. However, the mechanisms that control channel gating are unclear. Here we present crystal structures of the human TREK-2 channel (up to 3.4 angstrom resolution) in two conformations and in complex with norfluoxetine, the active metabolite of fluoxetine (Prozac) and a state-dependent blocker of TREK channels. Norfluoxetine binds within intramembrane fenestrations found in only one of these two conformations. Channel activation by arachidonic acid and mechanical stretch involves conversion between these states through movement of the pore-lining helices. These results provide an explanation for TREK channel mechanosensitivity, regulation by diverse stimuli, and possible off-target effects of the serotonin reuptake inhibitor Prozac.
- Structural Genomics Consortium, University of Oxford, Oxford OX3 7DQ, UK.
Organizational Affiliation: