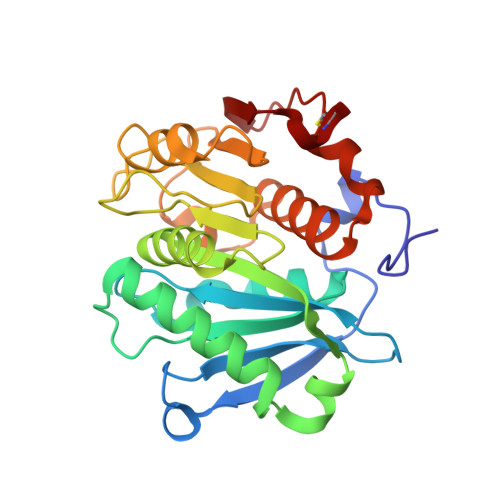
Crystal structure and thermodynamic and kinetic stability of metagenome-derived LC-cutinase.
Sulaiman, S., You, D.J., Kanaya, E., Koga, Y., Kanaya, S.(2014) Biochemistry 53: 1858-1869
- PubMed: 24593046 Search on PubMed
- DOI: https://doi.org/10.1021/bi401561p
- Primary Citation Related Structures:
4EB0 - PubMed Abstract:
The crystal structure of metagenome-derived LC-cutinase with polyethylene terephthalate (PET)-degrading activity was determined at 1.5 Å resolution. The structure strongly resembles that of Thermobifida alba cutinase. Ser165, Asp210, and His242 form the catalytic triad. Thermal denaturation and guanidine hydrochloride (GdnHCl)-induced unfolding of LC-cutinase were analyzed at pH 8.0 by circular dichroism spectroscopy. The midpoint of the transition of the thermal denaturation curve, T1/2, and that of the GdnHCl-induced unfolding curve, Cm, at 30 °C were 86.2 °C and 4.02 M, respectively. The free energy change of unfolding in the absence of GdnHCl, ΔG(H2O), was 41.8 kJ mol(-1) at 30 °C. LC-cutinase unfolded very slowly in GdnHCl with an unfolding rate, ku(H2O), of 3.28 × 10(-6) s(-1) at 50 °C. These results indicate that LC-cutinase is a kinetically robust protein. Nevertheless, the optimal temperature for the activity of LC-cutinase toward p-nitrophenyl butyrate (50 °C) was considerably lower than the T1/2 value. It increased by 10 °C in the presence of 1% polyethylene glycol (PEG) 1000. It also increased by at least 20 °C when PET was used as a substrate. These results suggest that the active site is protected from a heat-induced local conformational change by binding of PEG or PET. LC-cutinase contains one disulfide bond between Cys275 and Cys292. To examine whether this disulfide bond contributes to the thermodynamic and kinetic stability of LC-cutinase, C275/292A-cutinase without this disulfide bond was constructed. Thermal denaturation studies and equilibrium and kinetic studies of the GdnHCl-induced unfolding of C275/292A-cutinase indicate that this disulfide bond contributes not only to the thermodynamic stability but also to the kinetic stability of LC-cutinase.
- Department of Material and Life Science, Graduate School of Engineering, Osaka University , 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: