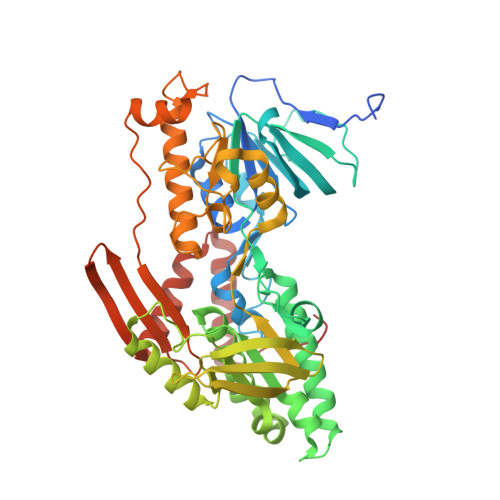
Structural insight into the type-II mitochondrial NADH dehydrogenases.
Feng, Y., Li, W., Li, J., Wang, J., Ge, J., Xu, D., Liu, Y., Wu, K., Zeng, Q., Wu, J.W., Tian, C., Zhou, B., Yang, M.(2012) Nature 491: 478-482
- PubMed: 23086143 Search on PubMed
- DOI: https://doi.org/10.1038/nature11541
- Primary Citation Related Structures:
4G6G, 4G6H, 4G73, 4G74 - PubMed Abstract:
The single-component type-II NADH dehydrogenases (NDH-2s) serve as alternatives to the multisubunit respiratory complex I (type-I NADH dehydrogenase (NDH-1), also called NADH:ubiquinone oxidoreductase; EC 1.6.5.3) in catalysing electron transfer from NADH to ubiquinone in the mitochondrial respiratory chain. The yeast NDH-2 (Ndi1) oxidizes NADH on the matrix side and reduces ubiquinone to maintain mitochondrial NADH/NAD(+) homeostasis. Ndi1 is a potential therapeutic agent for human diseases caused by complex I defects, particularly Parkinson's disease, because its expression restores the mitochondrial activity in animals with complex I deficiency. NDH-2s in pathogenic microorganisms are viable targets for new antibiotics. Here we solve the crystal structures of Ndi1 in its substrate-free, NADH-, ubiquinone- and NADH-ubiquinone-bound states, to help understand the catalytic mechanism of NDH-2s. We find that Ndi1 homodimerization through its carboxy-terminal domain is critical for its catalytic activity and membrane targeting. The structures reveal two ubiquinone-binding sites (UQ(I) and UQ(II)) in Ndi1. NADH and UQ(I) can bind to Ndi1 simultaneously to form a substrate-protein complex. We propose that UQ(I) interacts with FAD to act as an intermediate for electron transfer, and that NADH transfers electrons through this FAD-UQ(I) complex to UQ(II). Together our data reveal the regulatory and catalytic mechanisms of Ndi1 and may facilitate the development or targeting of NDH-2s for potential therapeutic applications.
- State Key Laboratory of Biomembrane and Membrane Biotechnology, Tsinghua-Peking Center for Life Sciences, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: