Crystal Structure of JMJD2A Complexed with Inhibitor
King, O.N.F., Krojer, T., Arrowsmith, C.H., Edwards, A., Bountra, C., McDonough, M.A., Schofield, C.J., Structural Genomics Consortium (SGC)To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Lysine-specific demethylase 4A | 381 | Homo sapiens | Mutation(s): 0 Gene Names: KDM4A, JHDM3A, JMJD2, JMJD2A, KIAA0677 EC: 1.14.11 (PDB Primary Data), 1.14.11.66 (UniProt), 1.14.11.69 (UniProt) | 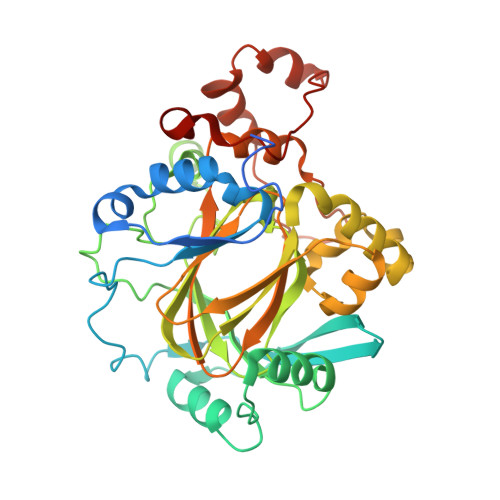 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O75164 GTEx: ENSG00000066135 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O75164 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 0WS Download:Ideal Coordinates CCD File | F [auth A], L [auth B] | 2-(1H-pyrazol-3-yl)pyridine-4-carboxylic acid C9 H7 N3 O2 NTNYOLXCVCDWFY-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | D [auth A], H [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| NI Download:Ideal Coordinates CCD File | C [auth A], G [auth B] | NICKEL (II) ION Ni VEQPNABPJHWNSG-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | E [auth A], I [auth B], J [auth B], K [auth B] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 100.75 | α = 90 |
| b = 150.03 | β = 90 |
| c = 57.49 | γ = 90 |
| Software Name | Purpose |
|---|---|
| GDA | data collection |
| PHASER | phasing |
| BUSTER | refinement |
| XDS | data reduction |
| SCALA | data scaling |