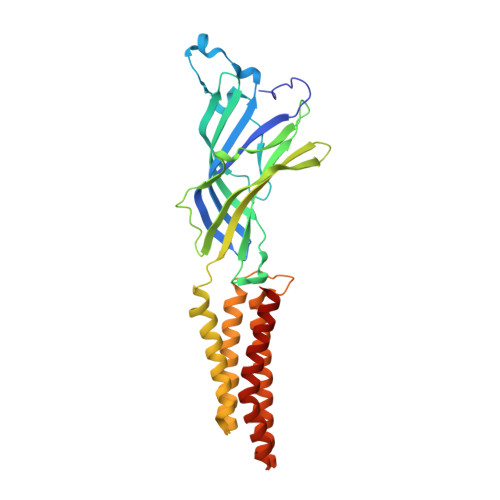
Structural basis for potentiation by alcohols and anaesthetics in a ligand-gated ion channel.
Sauguet, L., Howard, R.J., Malherbe, L., Lee, U.S., Corringer, P.J., Harris, R.A., Delarue, M.(2013) Nat Commun 4: 1697-1697
- PubMed: 23591864 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms2682
- Primary Citation Related Structures:
4HFB, 4HFC, 4HFD, 4HFE, 4HFH - PubMed Abstract:
Ethanol alters nerve signalling by interacting with proteins in the central nervous system, particularly pentameric ligand-gated ion channels. A recent series of mutagenesis experiments on Gloeobacter violaceus ligand-gated ion channel, a prokaryotic member of this family, identified a single-site variant that is potentiated by pharmacologically relevant concentrations of ethanol. Here we determine crystal structures of the ethanol-sensitized variant in the absence and presence of ethanol and related modulators, which bind in a transmembrane cavity between channel subunits and may stabilize the open form of the channel. Structural and mutagenesis studies defined overlapping mechanisms of potentiation by alcohols and anaesthetics via the inter-subunit cavity. Furthermore, homology modelling show this cavity to be conserved in human ethanol-sensitive glycine and GABA(A) receptors, and to involve residues previously shown to influence alcohol and anaesthetic action on these proteins. These results suggest a common structural basis for ethanol potentiation of an important class of targets for neurological actions of ethanol.
- Unité de Dynamique Structurale des Macromolécules, Institut Pasteur, F-75015 Paris, France.
Organizational Affiliation: