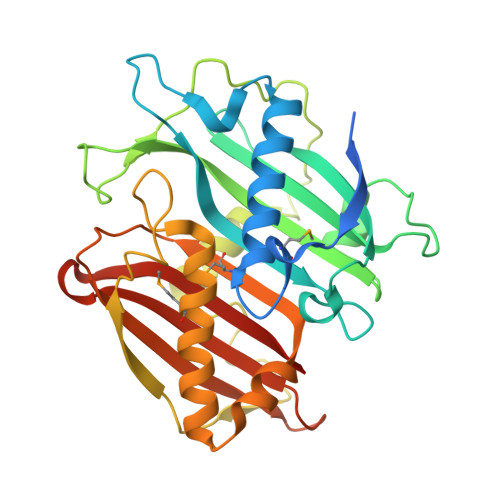
Structure of the Bifunctional Acyltransferase/Decarboxylase LnmK from the Leinamycin Biosynthetic Pathway Revealing Novel Activity for a Double-Hot-Dog Fold.
Lohman, J.R., Bingman, C.A., Phillips, G.N., Shen, B.(2013) Biochemistry 52: 902-911
- PubMed: 23320975 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi301652y
- Primary Citation Related Structures:
4HZN, 4HZO, 4HZP - PubMed Abstract:
The β-branched C3 unit in leinamycin biosynthesis is installed by a set of four proteins, LnmFKLM. In vitro biochemical investigation confirmed that LnmK is a bifunctional acyltransferase/decarboxylase (AT/DC) that catalyzes first self-acylation using methylmalonyl-CoA as a substrate and subsequently transacylation of the methylmalonyl group to the phosphopantetheinyl group of the LnmL acyl carrier protein [Liu, T., Huang, Y., and Shen, B. (2009) J. Am. Chem. Soc. 131, 6900-6901]. LnmK shows no sequence homology to proteins of known function, representing a new family of AT/DC enzymes. Here we report the X-ray structure of LnmK. LnmK is homodimer with each of the monomers adopting a double-hot-dog fold. Cocrystallization of LnmK with methylmalonyl-CoA revealed an active site tunnel terminated by residues from the dimer interface. In contrast to canonical AT and ketosynthase enzymes that employ Ser or Cys as an active site residue, none of these residues are found in the vicinity of the LnmK active site. Instead, three tyrosines were identified, one of which, Tyr62, was established, by site-directed mutagenesis, to be the most likely active site residue for the AT activity of LnmK. LnmK represents the first AT enzyme that employs a Tyr as an active site residue and the first member of the family of double-hot-dog fold enzymes that displays an AT activity known to date. The LnmK structure sets the stage for probing of the DC activity of LnmK through site-directed mutagenesis. These findings highlight natural product biosynthetic machinery as a rich source of novel enzyme activities, mechanisms, and structures.
- Department of Chemistry, Scripps Research Institute, The Scripps Research Institute, Jupiter, Florida 33485, United States.
Organizational Affiliation: