Enhanced Potency of a Broadly Neutralizing HIV-1 Antibody In Vitro Improves Protection against Lentiviral Infection In Vivo.
Rudicell, R.S., Kwon, Y.D., Ko, S.Y., Pegu, A., Louder, M.K., Georgiev, I.S., Wu, X., Zhu, J., Boyington, J.C., Chen, X., Shi, W., Yang, Z.Y., Doria-Rose, N.A., McKee, K., O'Dell, S., Schmidt, S.D., Chuang, G.Y., Druz, A., Soto, C., Yang, Y., Zhang, B., Zhou, T., Todd, J.P., Lloyd, K.E., Eudailey, J., Roberts, K.E., Donald, B.R., Bailer, R.T., Ledgerwood, J., Mullikin, J.C., Shapiro, L., Koup, R.A., Graham, B.S., Nason, M.C., Connors, M., Haynes, B.F., Rao, S.S., Roederer, M., Kwong, P.D., Mascola, J.R., Nabel, G.J.(2014) J Virol 88: 12669-12682
- PubMed: 25142607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02213-14
- Primary Citation Related Structures:
4OLU, 4OLV, 4OLW, 4OLX, 4OLY, 4OLZ, 4OM0, 4OM1 - PubMed Abstract:
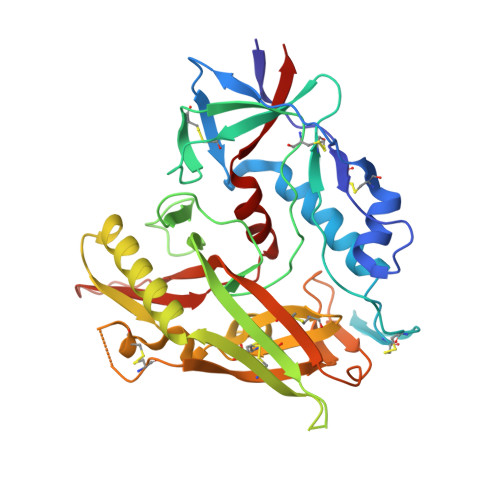
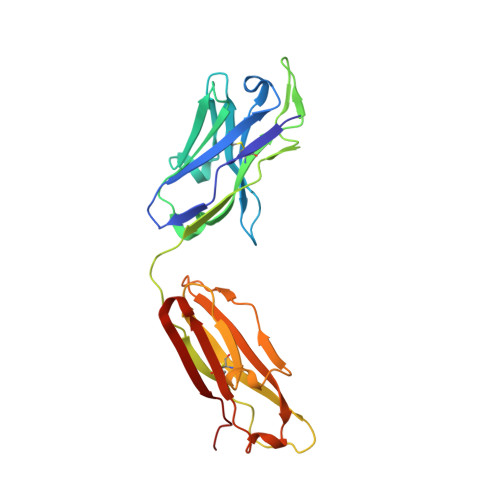
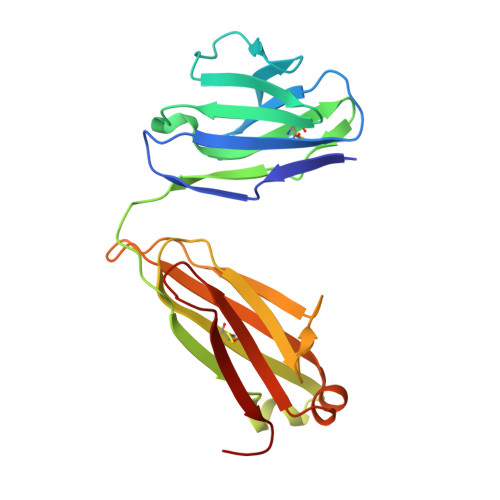
Over the past 5 years, a new generation of highly potent and broadly neutralizing HIV-1 antibodies has been identified. These antibodies can protect against lentiviral infection in nonhuman primates (NHPs), suggesting that passive antibody transfer would prevent HIV-1 transmission in humans. To increase the protective efficacy of such monoclonal antibodies, we employed next-generation sequencing, computational bioinformatics, and structure-guided design to enhance the neutralization potency and breadth of VRC01, an antibody that targets the CD4 binding site of the HIV-1 envelope. One variant, VRC07-523, was 5- to 8-fold more potent than VRC01, neutralized 96% of viruses tested, and displayed minimal autoreactivity. To compare its protective efficacy to that of VRC01 in vivo, we performed a series of simian-human immunodeficiency virus (SHIV) challenge experiments in nonhuman primates and calculated the doses of VRC07-523 and VRC01 that provide 50% protection (EC50). VRC07-523 prevented infection in NHPs at a 5-fold lower concentration than VRC01. These results suggest that increased neutralization potency in vitro correlates with improved protection against infection in vivo, documenting the improved functional efficacy of VRC07-523 and its potential clinical relevance for protecting against HIV-1 infection in humans. In the absence of an effective HIV-1 vaccine, alternative strategies are needed to block HIV-1 transmission. Direct administration of HIV-1-neutralizing antibodies may be able to prevent HIV-1 infections in humans. This approach could be especially useful in individuals at high risk for contracting HIV-1 and could be used together with antiretroviral drugs to prevent infection. To optimize the chance of success, such antibodies can be modified to improve their potency, breadth, and in vivo half-life. Here, knowledge of the structure of a potent neutralizing antibody, VRC01, that targets the CD4-binding site of the HIV-1 envelope protein was used to engineer a next-generation antibody with 5- to 8-fold increased potency in vitro. When administered to nonhuman primates, this antibody conferred protection at a 5-fold lower concentration than the original antibody. Our studies demonstrate an important correlation between in vitro assays used to evaluate the therapeutic potential of antibodies and their in vivo effectiveness.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, Maryland, USA.
Organizational Affiliation: