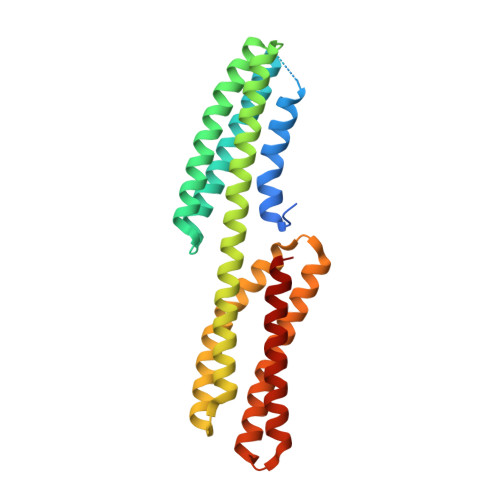
Structure of the free form of the N-terminal VH1 domain of monomeric alpha-catenin.
Shibahara, T., Hirano, Y., Hakoshima, T.(2015) FEBS Lett 589: 1754-1760
- PubMed: 26071377 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2015.05.053
- Primary Citation Related Structures:
4P9T - PubMed Abstract:
The N-terminal vinculin-homology 1 (VH1) domain of α-catenin facilitates two exclusive forms, a monomeric form directly bound to β-catenin for linking E-cadherin to F-actin or a homodimer for the inhibition of β-catenin binding. Competition of these two forms is affected by ∼80 N-terminal residues, whose structure is poorly understood. We have determined the structure of the monomeric free form of the αN-catenin VH1 domain and revealed that the N-terminal residues form α1 and α2 helices to complete formation of the N-terminal four-helix bundle. Dynamic conformational changes of these two helices control formation of the β-catenin-bound monomer or unbound homodimer.
- Structural Biology Laboratory, Nara Institute of Science and Technology, 8916-5 Takayama, Ikoma, Nara 630-0192, Japan.
Organizational Affiliation: