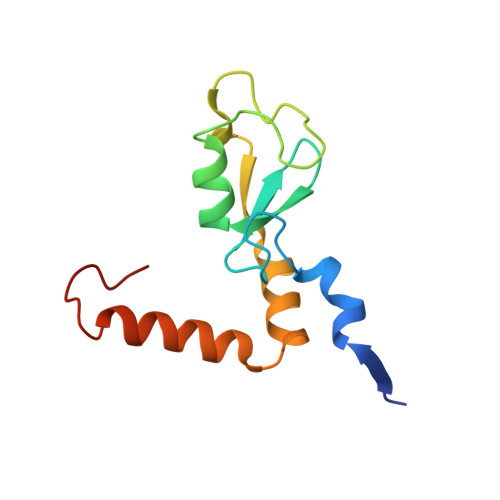
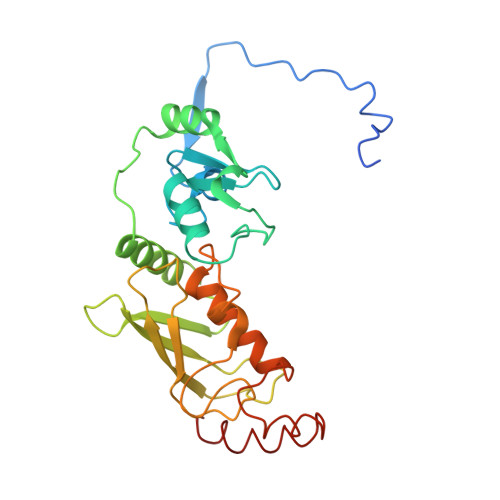
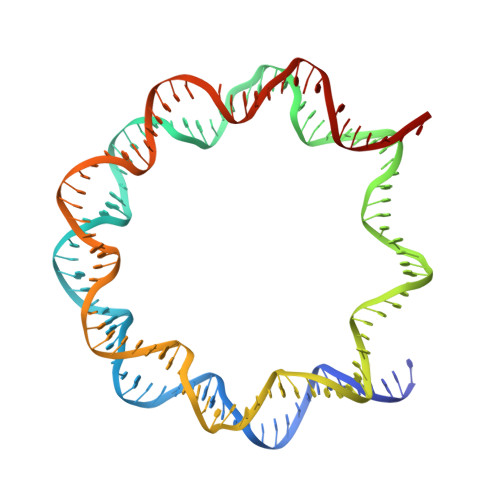
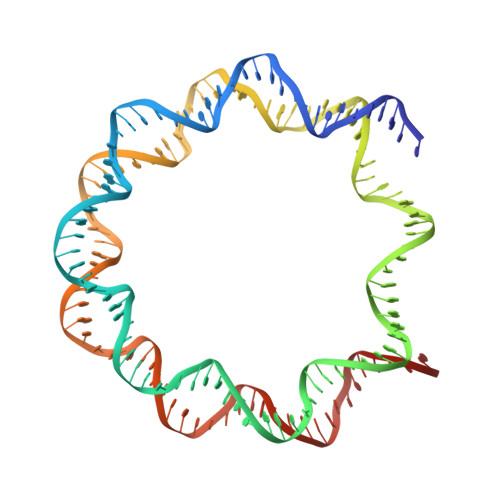
Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome.
McGinty, R.K., Henrici, R.C., Tan, S.(2014) Nature 514: 591-596
- PubMed: 25355358 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13890
- Primary Citation Related Structures:
4R8P - PubMed Abstract:
The Polycomb group of epigenetic enzymes represses expression of developmentally regulated genes in many eukaryotes. This group includes the Polycomb repressive complex 1 (PRC1), which ubiquitylates nucleosomal histone H2A Lys 119 using its E3 ubiquitin ligase subunits, Ring1B and Bmi1, together with an E2 ubiquitin-conjugating enzyme, UbcH5c. However, the molecular mechanism of nucleosome substrate recognition by PRC1 or other chromatin enzymes is unclear. Here we present the crystal structure of the human Ring1B-Bmi1-UbcH5c E3-E2 complex (the PRC1 ubiquitylation module) bound to its nucleosome core particle substrate. The structure shows how a chromatin enzyme achieves substrate specificity by interacting with several nucleosome surfaces spatially distinct from the site of catalysis. Our structure further reveals an unexpected role for the ubiquitin E2 enzyme in substrate recognition, and provides insight into how the related histone H2A E3 ligase, BRCA1, interacts with and ubiquitylates the nucleosome.
- Center for Eukaryotic Gene Regulation, Department of Biochemistry and Molecular Biology, The Pennsylvania State University, University Park, Pennsylvania 16802, USA.
Organizational Affiliation: