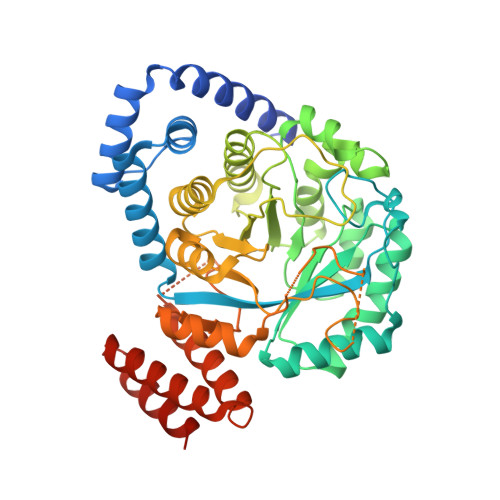
Crystal Structure of HydG from Carboxydothermus hydrogenoformans: A Trifunctional [FeFe]-Hydrogenase Maturase.
Nicolet, Y., Pagnier, A., Zeppieri, L., Martin, L., Amara, P., Fontecilla-Camps, J.C.(2015) Chembiochem 16: 397-402
- PubMed: 25504963 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201402661
- Primary Citation Related Structures:
4RTB - PubMed Abstract:
The structure of the radical S-adenosyl-L-methionine (SAM) [FeFe]-hydrogenase maturase HydG involved in CN(-) /CO synthesis is characterized by two internal tunnels connecting its tyrosine-binding pocket with the external medium and the C-terminal Fe4 S4 cluster-containing region. A comparison with a tryptophan-bound NosL structure suggests that substrate binding causes the closing of the first tunnel and, along with mutagenesis studies, that tyrosine binds to HydG with its amino group well positioned for H-abstraction by SAM. In this orientation the dehydroglycine (DHG) fragment caused by tyrosine Cα-Cβ bond scission can readily migrate through the second tunnel towards the C-terminal domain where both CN(-) and CO are synthesized. Our HydG structure appears to be in a relaxed state with its C-terminal cluster CysX2 CysX22 Cys motif exposed to solvent. A rotation of this domain coupled to Fe4 S4 cluster assembly would bury its putatively reactive unique Fe ion thereby allowing it to interact with DHG.
- Metalloproteins Unit, Institut de Biologie Structurale, UMR5075, CEA, CNRS, Université Grenoble Alpes, 71, Avenue des Martyrs, CS 10090, 38044 Grenoble cedex 9 (France). yvain.nicolet@ibs.fr.
Organizational Affiliation: