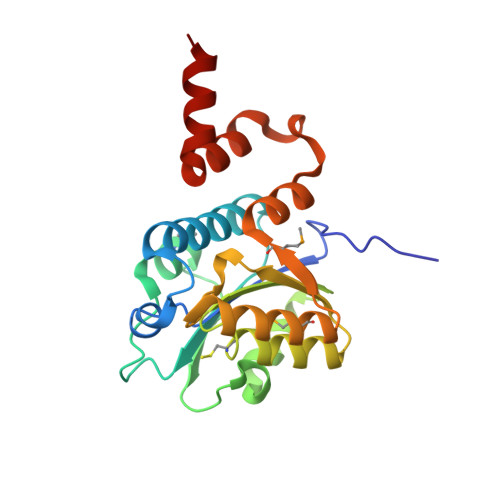
Crystal Structure of a Hidden Protein, YcaC, a Putative Cysteine Hydrolase from Pseudomonas aeruginosa, with and without an Acrylamide Adduct.
Grftehauge, M.K., Truan, D., Vasil, A., Denny, P.W., Vasil, M.L., Pohl, E.(2015) Int J Mol Sci 16: 15971-15984
- PubMed: 26184183 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms160715971
- Primary Citation Related Structures:
4WGF, 4WH0 - PubMed Abstract:
As part of the ongoing effort to functionally and structurally characterize virulence factors in the opportunistic pathogen Pseudomonas aeruginosa, we determined the crystal structure of YcaC co-purified with the target protein at resolutions of 2.34 and 2.56 Å without a priori knowledge of the protein identity or experimental phases. The three-dimensional structure of YcaC adopts a well-known cysteine hydrolase fold with the putative active site residues conserved. The active site cysteine is covalently bound to propionamide in one crystal form, whereas the second form contains an S-mercaptocysteine. The precise biological function of YcaC is unknown; however, related prokaryotic proteins have functions in antibacterial resistance, siderophore production and NADH biosynthesis. Here, we show that YcaC is exceptionally well conserved across both bacterial and fungal species despite being non-ubiquitous. This suggests that whilst YcaC may not be part of an integral pathway, the function could confer a significant evolutionary advantage to microbial life.
- School of Biological and Biomedical Sciences, Durham University, Durham DH1 3LE, UK. m.k.groftehauge@durham.ac.uk.
Organizational Affiliation: