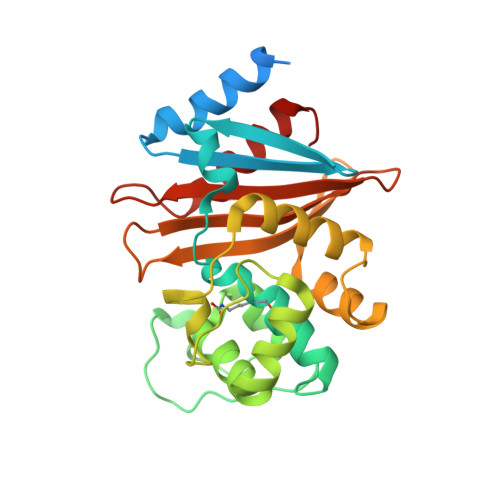
Structural Basis for Enhancement of Carbapenemase Activity in the OXA-51 Family of Class D beta-Lactamases.
Smith, C.A., Antunes, N.T., Stewart, N.K., Frase, H., Toth, M., Kantardjieff, K.A., Vakulenko, S.(2015) ACS Chem Biol 10: 1791-1796
- PubMed: 26042471
- DOI: https://doi.org/10.1021/acschembio.5b00090
- Primary Citation of Related Structures:
4ZDX - PubMed Abstract:
Class D β-lactamases of Acinetobacter baumannii are enzymes of the utmost clinical importance, producing resistance to last resort carbapenem antibiotics. Although the OXA-51-like enzymes constitute the largest family of class D β-lactamases, they are poorly studied and their importance in conferring carbapenem resistance is controversial. We present the detailed microbiological, kinetic, and structural characterization of the eponymous OXA-51 β-lactamase. Kinetic studies show that OXA-51 has low catalytic efficiency for carbapenems, primarily due to the low affinity of the enzyme for these substrates. Structural studies demonstrate that this low affinity results from the obstruction of the enzyme active site by the side chain of Trp222, which presents a transient steric barrier to an incoming carbapenem substrate. The Trp222Met substitution relieves this steric hindrance and elevates the affinity of the mutant enzyme for carbapenems by 10-fold, significantly increasing the levels of resistance to these antibiotics. The ability of OXA-51 to evolve into a robust carbapenemase as the result of a single amino acid substitution may, in the near future, elevate the ubiquitous enzymes of the OXA-51 family to the status of the most deleterious A. baumannii carbapenemases, with dire clinical consequences.
- †Stanford Synchrotron Radiation Lightsource, Stanford University, Menlo Park, California 94025, United States.
Organizational Affiliation: