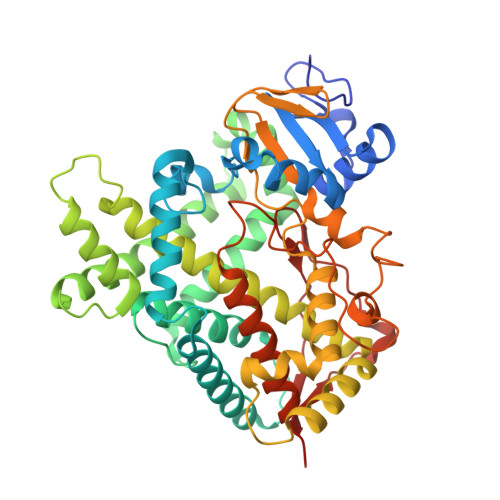
Coumarin Derivatives as Substrate Probes of Mammalian Cytochromes P450 2B4 and 2B6: Assessing the Importance of 7-Alkoxy Chain Length, Halogen Substitution, and Non-Active Site Mutations.
Liu, J., Shah, M.B., Zhang, Q., Stout, C.D., Halpert, J.R., Wilderman, P.R.(2016) Biochemistry 55: 1997-2007
- PubMed: 26982502 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.5b01330
- Primary Citation Related Structures:
4ZV8, 5EM4 - PubMed Abstract:
Using a combined structural and biochemical approach, the functional importance of a recently described peripheral pocket bounded by the E-, F-, G-, and I-helices in CYP2B4 and 2B6 was probed. Three series of 4-substituted-7-alkoxycoumarin derivatives with -H, -CH3, or -CF3 at the 4 position of the coumarin core were used initially to monitor functional differences between CYP2B4 and 2B6. 7-Ethoxy-4-(trifluoromethyl)coumarin (7-EFC) displayed the highest catalytic efficiency among these substrates. Mutants were made to alter side-chain polarity (V/E194Q) or bulk (F/Y244W) to alter access to the peripheral pocket. Modest increases in catalytic efficiency of 7-EFC O-deethylation by the mutants were magnified considerably by chlorination or bromination of the substrate ethoxy chain. A structure of CYP2B6 Y244W in complex with (+)-α-pinene was solved at 2.2 Å and showed no CYMAL-5 in the peripheral pocket. A ligand free structure of CYP2B4 F244W was solved at 3.0 Å with CYMAL-5 in the peripheral pocket. In both instances, comparison of the respective wild-type and mutant CYP2B enzymes revealed that CYMAL-5 occupancy of the peripheral pocket had little effect on the topology of active site residue side-chains, despite the fact that the peripheral pocket and active site are located on opposite sides of the I-helix. Analysis of available CYP2B structures suggest that the effect of the amino acid substitutions within the peripheral pocket derive from altered interactions between the F and G helices.
- School of Pharmacy, University of Connecticut , Storrs, Connecticut 06269, United States.
Organizational Affiliation: