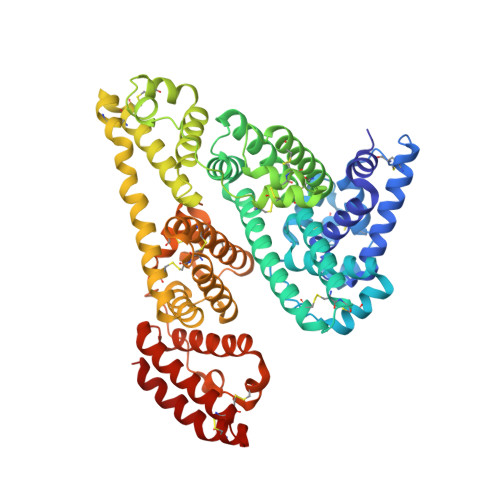
Structural studies of bovine, equine, and leporine serum albumin complexes with naproxen.
Bujacz, A., Zielinski, K., Sekula, B.(2014) Proteins 82: 2199-2208
- PubMed: 24753230 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24583
- Primary Citation Related Structures:
4OR0, 4OT2, 4PO0 - PubMed Abstract:
Serum albumin, a protein naturally abundant in blood plasma, shows remarkable ligand binding properties of numerous endogenous and exogenous compounds. Most of serum albumin binding sites are able to interact with more than one class of ligands. Determining the protein-ligand interactions among mammalian serum albumins is essential for understanding the complexity of this transporter. We present three crystal structures of serum albumins in complexes with naproxen (NPS): bovine (BSA-NPS), equine (ESA-NPS), and leporine (LSA-NPS) determined to 2.58 Å (C2), 2.42 Å (P61), and 2.73 Å (P2₁2₁2₁) resolutions, respectively. A comparison of the structurally investigated complexes with the analogous complex of human serum albumin (HSA-NPS) revealed surprising differences in the number and distribution of naproxen binding sites. Bovine and leporine serum albumins possess three NPS binding sites, but ESA has only two. All three complexes of albumins studied here have two common naproxen locations, but BSA and LSA differ in the third NPS binding site. None of these binding sites coincides with the naproxen location in the HSA-NPS complex, which was obtained in the presence of other ligands besides naproxen. Even small differences in sequences of serum albumins from various species, especially in the area of the binding pockets, influence the affinity and the binding mode of naproxen to this transport protein.
- Institute of Technical Biochemistry, Faculty of Biotechnology and Food Sciences, Lodz University of Technology, Stefanowskiego 4/10, 90-924, Lodz, Poland.
Organizational Affiliation: