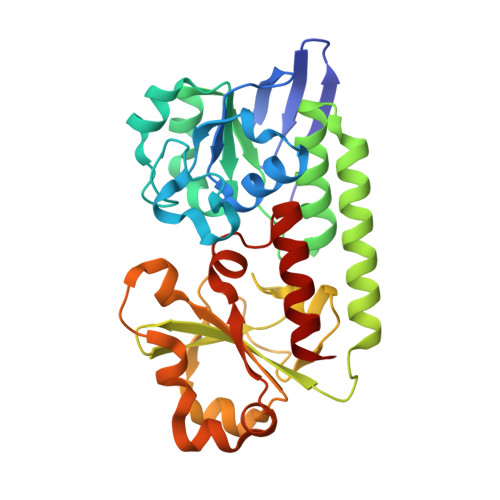
Interactions of a Periplasmic Binding Protein with a Tetradentate Siderophore Mimic.
Raines, D.J., Moroz, O.V., Wilson, K.S., Duhme-Klair, A.(2013) Angew Chem Int Ed Engl 52: 4595
- PubMed: 23512642 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201300751
- Primary Citation Related Structures:
3ZKW, 5A1J - PubMed Abstract:
Iron-bound structure: The ferric complex of a tetradentate siderophore mimic was synthesized and co-crystallized with the periplasmic binding protein CeuE of Campylobacter jejuni. In addition to electrostatic and hydrogen-bonding interactions between the binding pocket and the substrate, the structure showed direct coordination of two amino acid side chains to the Fe(III) center (orange, see figure).
- Department of Chemistry, University of York, Heslington, York, YO10 5DD, UK.
Organizational Affiliation: