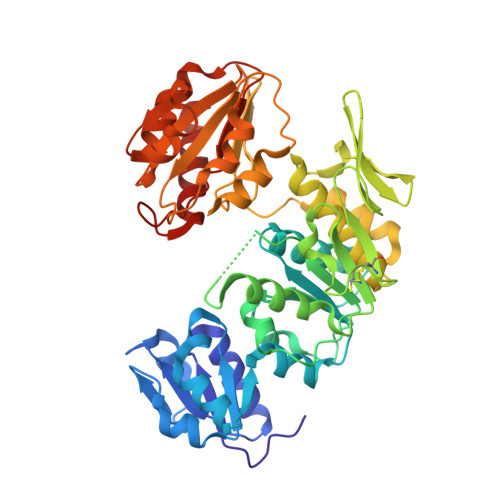
Crystallographic Study of Peptidoglycan Biosynthesis Enzyme MurD: Domain Movement Revisited.
Sink, R., Kotnik, M., Zega, A., Barreteau, H., Gobec, S., Blanot, D., Dessen, A., Contreras-Martel, C.(2016) PLoS One 11: e0152075-e0152075
- PubMed: 27031227 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0152075
- Primary Citation Related Structures:
5A5E, 5A5F - PubMed Abstract:
The biosynthetic pathway of peptidoglycan, an essential component of bacterial cell wall, is a well-recognized target for antibiotic development. Peptidoglycan precursors are synthesized in the bacterial cytosol by various enzymes including the ATP-hydrolyzing Mur ligases, which catalyze the stepwise addition of amino acids to a UDP-MurNAc precursor to yield UDP-MurNAc-pentapeptide. MurD catalyzes the addition of D-glutamic acid to UDP-MurNAc-L-Ala in the presence of ATP; structural and biochemical studies have suggested the binding of the substrates with an ordered kinetic mechanism in which ligand binding inevitably closes the active site. In this work, we challenge this assumption by reporting the crystal structures of intermediate forms of MurD either in the absence of ligands or in the presence of small molecules. A detailed analysis provides insight into the events that lead to the closure of MurD and reveals that minor structural modifications contribute to major overall conformation alterations. These novel insights will be instrumental in the development of new potential antibiotics designed to target the peptidoglycan biosynthetic pathway.
- University of Ljubljana, Faculty of Pharmacy, Aškerčeva 7, Ljubljana, Slovenia.
Organizational Affiliation: