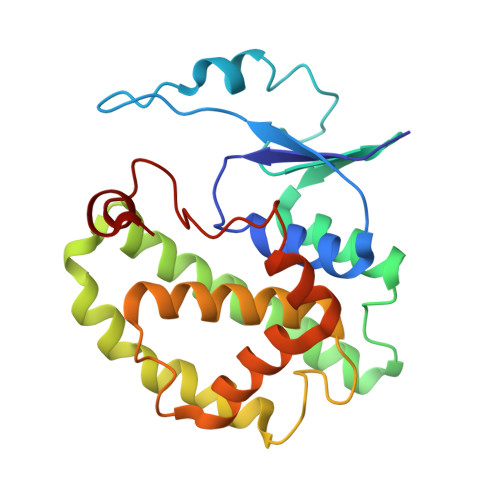
Crystal structure of a class-mu glutathione S-transferase from whiteleg shrimp Litopenaeus vannamei: structural changes in the xenobiotic binding H-site may alter the spectra of molecules bound.
Juarez-Martinez, A.B., Sotelo-Mundo, R.R., Rudino-Pinera, E.(2017) J Biochem Mol Toxicol 31
- PubMed: 27717103 Search on PubMed
- DOI: https://doi.org/10.1002/jbt.21838
- Primary Citation Related Structures:
5AN1 - PubMed Abstract:
Glutathione S-transferases (GSTs) are dimeric proteins that play a key role in phase II cellular detoxification. Here, the first crystal structure of a GST class-mu from marine crustacean shrimp Litopenaeus vannamei is reported at a resolution of 2.0 Å. The coordinates reported here have the lowest sequence identity with previously reported GSTs class-mu deposited at the Protein Data Bank (PDB), although they have subtle conformational differences. One key feature of GST class-mu from L. vannamei is the active site crevice markedly reduced when it is compared with other GSTs class-mu. This finding together with the chemical change of residues into the cavity (F112 and Y210) points to a particular specialization in which smallest xenobiotics with nonstandard chemical characteristics can be bound to the H-site. This suggests that marine organisms have evolved structural strategies to provide efficient selectivity toward xenobiotics to be disposed of by the phase II detoxification process.
- Departamento de Medicina Molecular y Bioprocesos, Instituto de Biotecnología, Universidad Nacional Autónoma de México, Colonia Chamilpa, 62210, Cuernavaca, Morelos, México.
Organizational Affiliation: