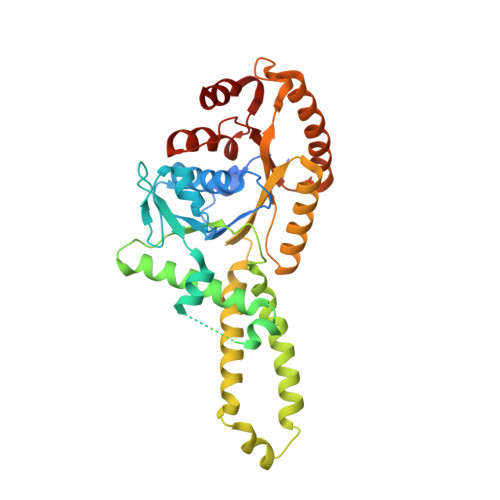
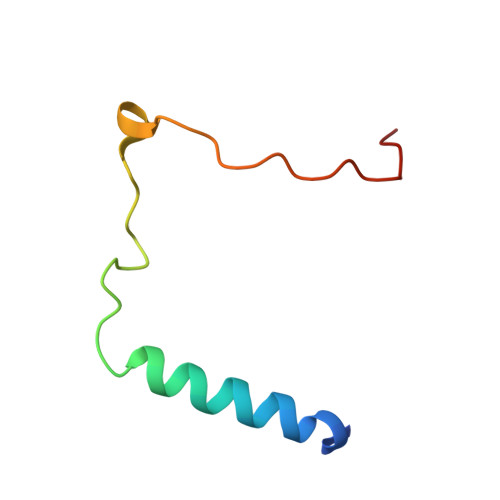
Mechanism of Assembly of a Substrate Transfer Complex during Tail-anchored Protein Targeting.
Gristick, H.B., Rome, M.E., Chartron, J.W., Rao, M., Hess, S., Shan, S.O., Clemons, W.M.(2015) J Biological Chem 290: 30006-30017
- PubMed: 26451041 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.677328
- Primary Citation Related Structures:
5BW8, 5BWK - PubMed Abstract:
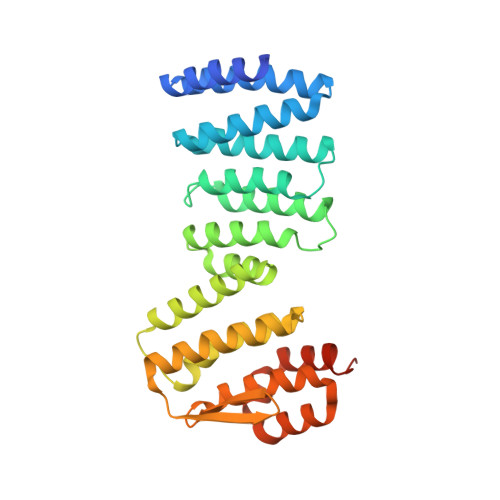
Tail-anchored (TA) proteins, defined as having a single transmembrane helix at their C terminus, are post-translationally targeted to the endoplasmic reticulum membrane by the guided entry of TA proteins (GET) pathway. In yeast, the handover of TA substrates is mediated by the heterotetrameric Get4/Get5 complex (Get4/5), which tethers the co-chaperone Sgt2 to the targeting factor, the Get3 ATPase. Binding of Get4/5 to Get3 is critical for efficient TA targeting; however, questions remain about the formation of the Get3·Get4/5 complex. Here we report crystal structures of a Get3·Get4/5 complex from Saccharomyces cerevisiae at 2.8 and 6.0 Å that reveal a novel interface between Get3 and Get4 dominated by electrostatic interactions. Kinetic and mutational analyses strongly suggest that these structures represent an on-pathway intermediate that rapidly assembles and then rearranges to the final Get3·Get4/5 complex. Furthermore, we provide evidence that the Get3·Get4/5 complex is dominated by a single Get4/5 heterotetramer bound to one monomer of a Get3 dimer, uncovering an intriguing asymmetry in the Get4/5 heterotetramer upon Get3 binding. Ultrafast diffusion-limited electrostatically driven Get3·Get4/5 association enables Get4/5 to rapidly sample and capture Get3 at different stages of the GET pathway.
- From the Division of Chemistry and Chemical Engineering and.
Organizational Affiliation: