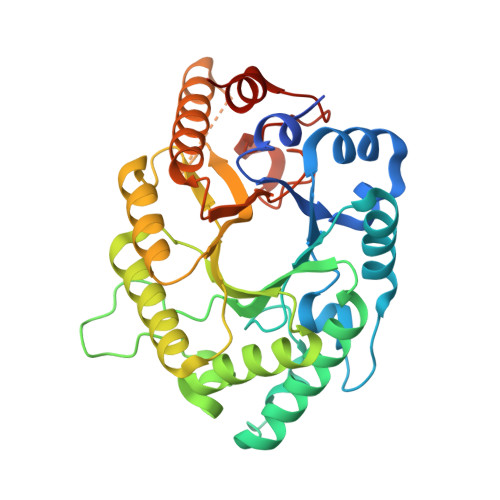
Structural insight into potential cold adaptation mechanism through a psychrophilic glycoside hydrolase family 10 endo-beta-1,4-xylanase.
Zheng, Y., Li, Y., Liu, W., Chen, C.C., Ko, T.P., He, M., Xu, Z., Liu, M., Luo, H., Guo, R.T., Yao, B., Ma, Y.(2016) J Struct Biol 193: 206-211
- PubMed: 26719223 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.12.010
- Primary Citation Related Structures:
5AY7, 5D4Y - PubMed Abstract:
The cold-adapted xylanases can catalyze at low temperature and hold great potential in food industry applications. Here we describe the first crystal structure of a cold-adapted glycoside hydrolase (GH) family 10 xylanase XynGR40 and its complex with xylobiose at 2.15 and 2.50Å resolution. The enzyme folds into a typical GH10 (β/α)8 TIM-barrel, with E132 and E243 serving as the catalytic residues. The xylobiose was observed to occupy the -1 and -2 subsites. Structural comparison with a thermophilic GH10 xylanase highlighting various parameters that may explain the cold adaptation features were analyzed. Synergistic effects of the increased exposure of hydrophobic residues, the higher flexibility of substrate-binding residues, more flexible loops, and the ratios of special amino acid residues, may result in the cold adaptation of XynGR40.
- Industrial Enzymes National Engineering Laboratory, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.
Organizational Affiliation: