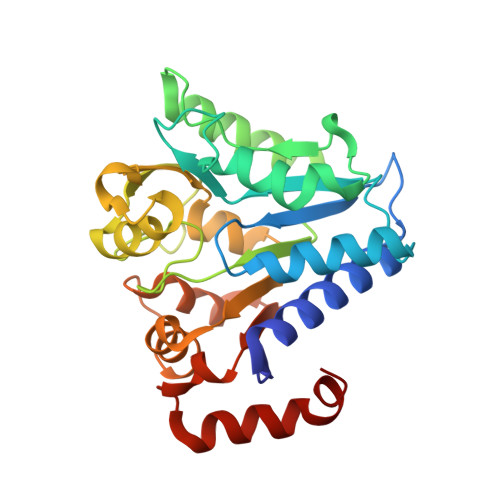
Crystal structure of tertiary complex of glucosamine-6-phosphate deaminase from Vibrio cholerae with BETA-D-GLUCOSE-6-PHOSPHATE and FRUCTOSE -6-PHOSPHATE
Chang, C., Maltseva, N., Kim, Y., Kwon, K., Anderson, W.F., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.