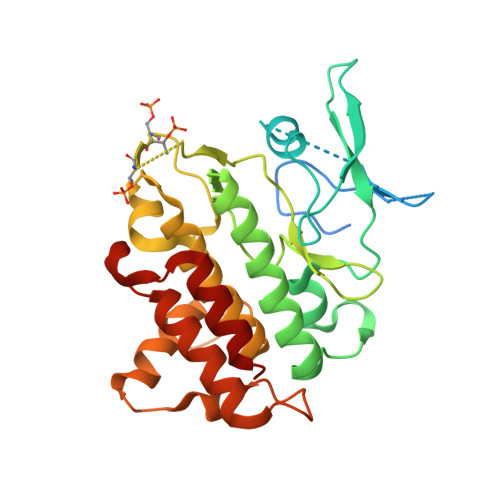
Efforts towards the optimization of a bi-aryl class of potent IRAK4 inhibitors.
Hanisak, J., Seganish, W.M., McElroy, W.T., Tang, H., Zhang, R., Tsui, H.C., Fischmann, T., Tulshian, D., Tata, J., Sondey, C., Devito, K., Fossetta, J., Garlisi, C.G., Lundell, D., Niu, X.(2016) Bioorg Med Chem Lett 26: 4250-4255
- PubMed: 27476420 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2016.07.048
- Primary Citation Related Structures:
5KX7, 5KX8 - PubMed Abstract:
IRAK4 has been identified as potential therapeutic target for inflammatory and autoimmune diseases. Herein we report the identification and initial SAR studies of a new class of pyrazole containing IRAK4 inhibitors designed to expand chemical diversity and improve off target activity of a previously identified series. These compounds maintain potent IRAK4 activity and desirable ligand efficiency. Rat clearance and a variety of off target activities were also examined, resulting in encouraging data with tractable SAR.
- Discovery Chemistry, Merck Research Laboratories, 2015 Galloping Hill Rd., Kenilworth, NJ 07033, Unites States. Electronic address: jennifer.hanisak@merck.com.
Organizational Affiliation: