Functional and structural analysis of AT-specific minor groove binders that disrupt DNA-protein interactions and cause disintegration of the Trypanosoma brucei kinetoplast.
Millan, C.R., Acosta-Reyes, F.J., Lagartera, L., Ebiloma, G.U., Lemgruber, L., Nue Martinez, J.J., Saperas, N., Dardonville, C., de Koning, H.P., Campos, J.L.(2017) Nucleic Acids Res 45: 8378-8391
- PubMed: 28637278 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx521
- Primary Citation Related Structures:
5LIT, 6GIM - PubMed Abstract:
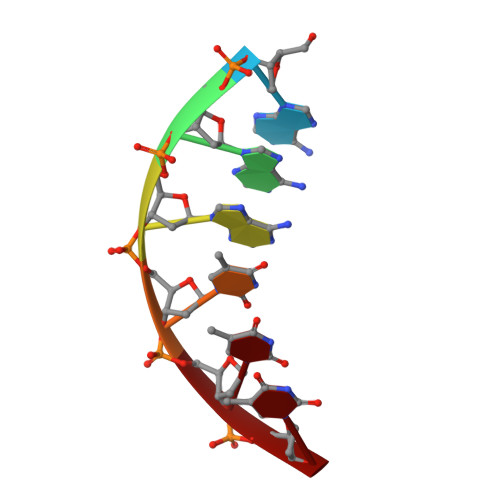
Trypanosoma brucei, the causative agent of sleeping sickness (Human African Trypanosomiasis, HAT), contains a kinetoplast with the mitochondrial DNA (kDNA), comprising of >70% AT base pairs. This has prompted studies of drugs interacting with AT-rich DNA, such as the N-phenylbenzamide bis(2-aminoimidazoline) derivatives 1 [4-((4,5-dihydro-1H-imidazol-2-yl)amino)-N-(4-((4,5-dihydro-1H-imidazol-2-yl)amino)phenyl)benzamide dihydrochloride] and 2 [N-(3-chloro-4-((4,5-dihydro-1H-imidazol-2-yl)amino)phenyl)-4-((4,5-dihydro-1H-imidazol-2-yl)amino)benzamide] as potential drugs for HAT. Both compounds show in vitro effects against T. brucei and in vivo curative activity in a mouse model of HAT. The main objective was to identify their cellular target inside the parasite. We were able to demonstrate that the compounds have a clear effect on the S-phase of T. brucei cell cycle by inflicting specific damage on the kinetoplast. Surface plasmon resonance (SPR)-biosensor experiments show that the drug can displace HMG box-containing proteins essential for kDNA function from their kDNA binding sites. The crystal structure of the complex of the oligonucleotide d[AAATTT]2 with compound 1 solved at 1.25 Å (PDB-ID: 5LIT) shows that the drug covers the minor groove of DNA, displaces bound water and interacts with neighbouring DNA molecules as a cross-linking agent. We conclude that 1 and 2 are powerful trypanocides that act directly on the kinetoplast, a structure unique to the order Kinetoplastida.
- Departament d'Enginyeria Química, EEBE, Universitat Politècnica de Catalunya, 08019 Barcelona, Spain.
Organizational Affiliation: