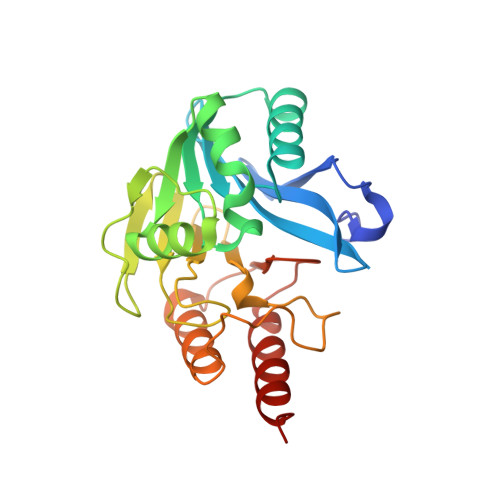
The structure of the metallo-beta-lactamase VIM-2 in complex with a triazolylthioacetamide inhibitor.
Christopeit, T., Yang, K.W., Yang, S.K., Leiros, H.K.(2016) Acta Crystallogr F Struct Biol Commun 72: 813-819
- PubMed: 27834790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16016113
- Primary Citation Related Structures:
5LSC - PubMed Abstract:
The increasing number of pathogens expressing metallo-β-lactamases (MBLs), and in this way achieving resistance to β-lactam antibiotics, is a significant threat to global public health. A promising strategy to treat such resistant pathogens is the co-administration of MBL inhibitors together with β-lactam antibiotics. However, an MBL inhibitor suitable for clinical use has not yet been identified. Verona integron-encoded metallo-β-lactamase 2 (VIM-2) is a widespread MBL with a broad substrate spectrum and hence is an interesting drug target for the treatment of β-lactam-resistant infections. In this study, three triazolylthioacetamides were tested as inhibitors of VIM-2. One of the tested compounds showed clear inhibition of VIM-2, with an IC 50 of 20 µM. The crystal structure of the inhibitor in complex with VIM-2 was obtained by DMSO-free co-crystallization and was solved at a resolution of 1.50 Å. To our knowledge, this is the first structure of a triazolylthioacetamide inhibitor in complex with an MBL. Analysis of the structure shows that the inhibitor binds to the two zinc ions in the active site of VIM-2 and revealed detailed information on the interactions involved. Furthermore, the crystal structure showed that binding of the inhibitor induced a conformational change of the conserved residue Trp87.
- The Norwegian Structural Biology Centre (Norstruct), Department of Chemistry, Faculty of Science and Technology, UiT The Arctic University of Norway, N-9037 Tromsø, Norway.
Organizational Affiliation: