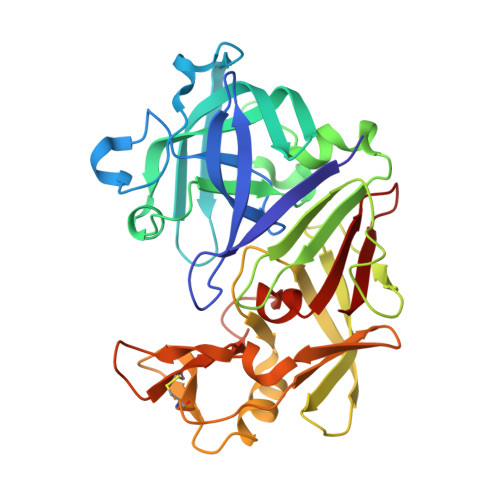
Design and Synthesis of Bioisosteres of Acylhydrazones as Stable Inhibitors of the Aspartic Protease Endothiapepsin.
Jumde, V.R., Mondal, M., Gierse, R.M., Unver, M.Y., Magari, F., van Lier, R.C.W., Heine, A., Klebe, G., Hirsch, A.K.H.(2018) ChemMedChem 13: 2266-2270
- PubMed: 30178575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.201800446
- Primary Citation Related Structures:
5OJE - PubMed Abstract:
Acylhydrazone-based dynamic combinatorial chemistry (DCC) is a powerful strategy for the rapid identification of novel hits. Even though acylhydrazones are important structural motifs in medicinal chemistry, their further progression in development may be hampered by major instability and potential toxicity under physiological conditions. It is therefore of paramount importance to identify stable replacements for acylhydrazone linkers. Herein, we present the first report on the design and synthesis of stable bioisosteres of acylhydrazone-based inhibitors of the aspartic protease endothiapepsin as a follow-up to a DCC study. The most successful bioisostere is equipotent, bears an amide linker, and we confirmed its binding mode by X-ray crystallography. Having some validated bioisosteres of acylhydrazones readily available might accelerate hit-to-lead optimization in future acylhydrazone-based DCC projects.
- Chemical Biology, Stratingh Institute for Chemistry, University of Groningen, Nijenborgh 7, 9747, AG, Groningen, The Netherlands.
Organizational Affiliation: