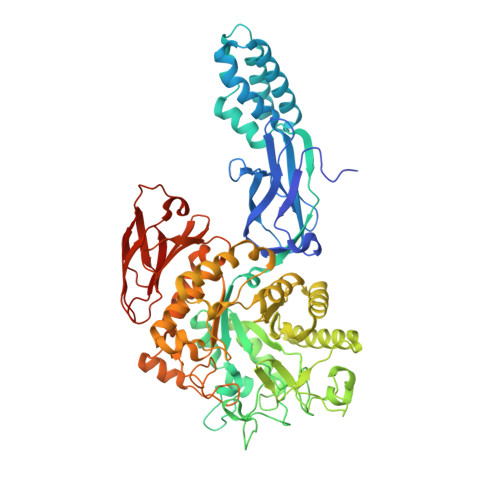
Zwitterionic pyrrolidene-phosphonates: inhibitors of the glycoside hydrolase-like phosphorylase Streptomyces coelicolor GlgEI-V279S.
Veleti, S.K., Petit, C., Ronning, D.R., Sucheck, S.J.(2017) Org Biomol Chem 15: 3884-3891
- PubMed: 28422240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c7ob00388a
- Primary Citation Related Structures:
5VSJ, 5VT4 - PubMed Abstract:
We synthesized and evaluated new zwitterionic inhibitors against glycoside hydrolase-like phosphorylase Streptomyces coelicolor (Sco) GlgEI-V279S which plays a role in α-glucan biosynthesis. Sco GlgEI-V279S serves as a model enzyme for validated anti-tuberculosis (TB) target Mycobacterium tuberculosis (Mtb) GlgE. Pyrrolidine inhibitors 5 and 6 were designed based on transition state considerations and incorporate a phosphonate on the pyrrolidine moiety to expand the interaction network between the inhibitor and the enzyme active site. Compounds 5 and 6 inhibited Sco GlgEI-V279S with K i = 45 ± 4 μM and 95 ± 16 μM, respectively, and crystal structures of Sco GlgE-V279S-5 and Sco GlgE-V279S-6 were obtained at a 3.2 Å and 2.5 Å resolution, respectively.
- Department of Chemistry and Biochemistry and School of Green Chemistry and Engineering, The University of Toledo, 2801 W. Bancroft Street, Toledo, Ohio 43606, USA. steve.sucheck@utoledo.edu donald.ronning@utoledo.edu.
Organizational Affiliation: