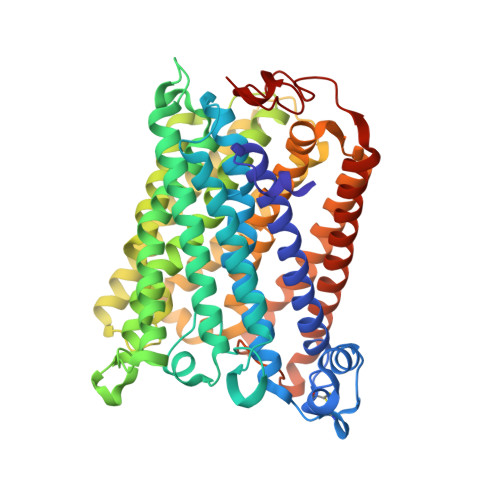
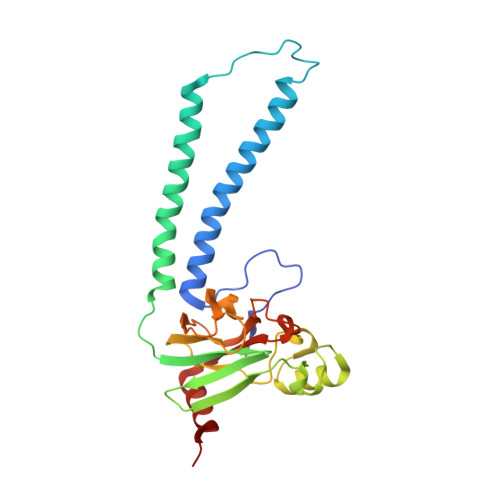
The K-path entrance in cytochrome c oxidase is defined by mutation of E101 and controlled by an adjacent ligand binding domain.
Hiser, C., Liu, J., Ferguson-Miller, S.(2018) Biochim Biophys Acta 1859: 725-733
- PubMed: 29626419 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbabio.2018.03.017
- Primary Citation Related Structures:
6CI0 - PubMed Abstract:
Three mutant forms of Rhodobacter sphaeroides cytochrome c oxidase (RsCcO) were created to test for multiple K-path entry sites (E101W), the existence of an "upper ligand site" (M350 W), and the nature and binding specificity of the "lower ligand site" (P315W/E101A) in the region of a crystallographically-defined deoxycholate at the K-path entrance. The effects of inhibitory and stimulatory detergents (dodecyl maltoside and Tween20) on these mutants are presented, as well as competition with other ligands, including the potentially physiologically relevant ligands cholesterol and retinoic acid. Ligands are shown to be able to compete with natural lipids to affect the activity of membrane-bound RsCcO. Results point to a single K-path entrance site at E101, with a single ligand binding pocket proximal to the entrance. The affinity of this pocket for amphipathic ligands is enhanced by removal of the E101 carboxyl and blocked by substituting a tryptophan in this area. A new crystal structure of the E101A mutant of RsCcO is presented that illustrates the structural basis of these results, showing that the loss of the E101 carboxyl creates a more hydrophobic groove consistent with altered ligand affinities.
- Department of Biochemistry and Molecular Biology, Michigan State University, East Lansing, MI 48824, United States. Electronic address: hiser@msu.edu.
Organizational Affiliation: