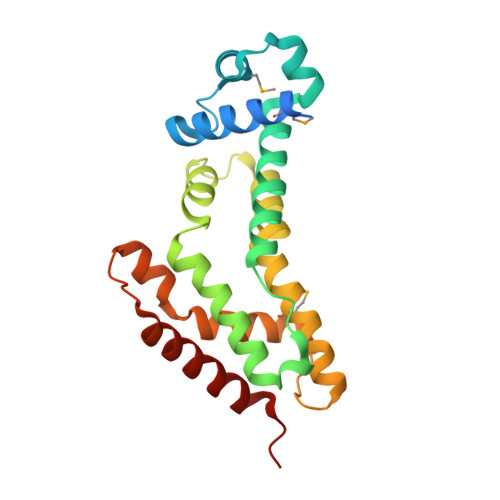
The induction mechanism of the flavonoid-responsive regulator FrrA.
Werner, N., Werten, S., Hoppen, J., Palm, G.J., Gottfert, M., Hinrichs, W.(2021) FEBS J
- PubMed: 34314575 Search on PubMed
- DOI: https://doi.org/10.1111/febs.16141
- Primary Citation Related Structures:
6G87, 6G8G, 6G8H - PubMed Abstract:
Bradyrhizobium diazoefficiens, a bacterial symbiont of soybean and other leguminous plants, enters a nodulation-promoting genetic programme in the presence of host-produced flavonoids and related signalling compounds. Here, we describe the crystal structure of an isoflavonoid-responsive regulator (FrrA) from Bradyrhizobium, as well as cocrystal structures with inducing and noninducing ligands (genistein and naringenin, respectively). The structures reveal a TetR-like fold whose DNA-binding domain is capable of adopting a range of orientations. A single molecule of either genistein or naringenin is asymmetrically bound in a central cavity of the FrrA homodimer, mainly via C-H contacts to the π-system of the ligands. Strikingly, however, the interaction does not provoke any conformational changes in the repressor. Both the flexible positioning of the DNA-binding domain and the absence of structural change upon ligand binding are corroborated by small-angle X-ray scattering (SAXS) experiments in solution. Together with a model of the promoter-bound state of FrrA our results suggest that inducers act as a wedge, preventing the DNA-binding domains from moving close enough together to interact with successive positions of the major groove of the palindromic operator.
- Institute for Biochemistry, Department Molecular Structural Biology, University of Greifswald, Germany.
Organizational Affiliation: