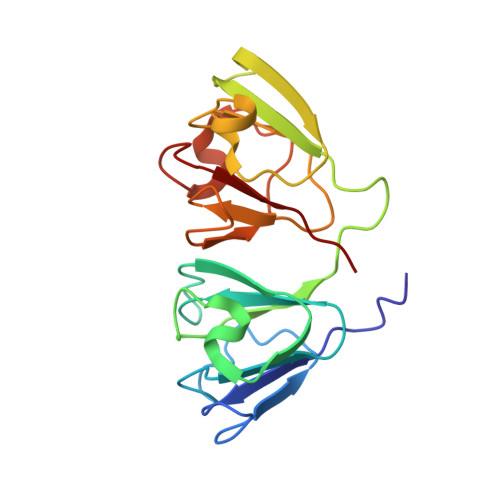
Structure of G57W mutant of human gamma S-crystallin and its involvement in cataract formation.
Bari, K.J., Sharma, S., Chary, K.V.R.(2019) J Struct Biol 205: 72-78
- PubMed: 30769148 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.02.003
- Primary Citation Related Structures:
6IF9 - PubMed Abstract:
A recently identified mutant of human γS-crystallin, G57W is associated with dominant congenital cataracts, the familial determinate of childhood blindness worldwide. To investigate the structural and functional changes that mediate the effect of this cataract-related mutant to compromise eye lens transparency and cause lens opacification in children, we recently reported complete sequence-specific resonance assignments of γS-G57W using a suite of heteronuclear NMR experiments. As a follow up, we have determined the 3D structure of γS-G57W and studied its conformational dynamics by solution NMR spectroscopy. Our structural dynamics results reveal greater flexibility of the N-terminal domain, which undergoes site-specific structural changes to accommodate W57, than its C-terminal counterpart. Our structural inferences that the unusual solvent exposure of W57 is associated with rearrangement of the N-terminal domain suggest an efficient pathway for increased aggregation in γS-G57W and illuminates the molecular dynamics underlying cataractogenic aggregation of lens crystallins in particular and aggregation of proteins in general.
- Center for Interdisciplinary Sciences, Tata Institute of Fundamental Research, Gopanpally, Hyderabad 500107, India.
Organizational Affiliation: