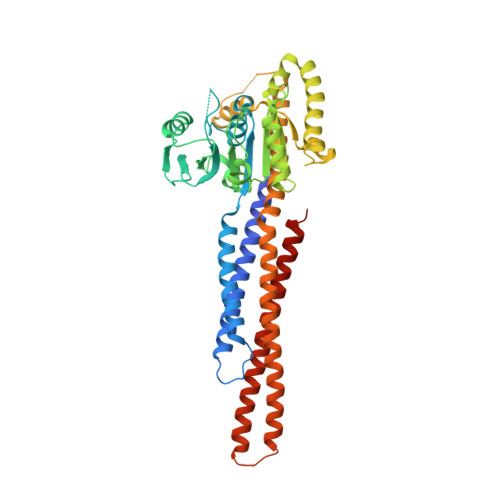
Structural insights of human mitofusin-2 into mitochondrial fusion and CMT2A onset.
Li, Y.J., Cao, Y.L., Feng, J.X., Qi, Y., Meng, S., Yang, J.F., Zhong, Y.T., Kang, S., Chen, X., Lan, L., Luo, L., Yu, B., Chen, S., Chan, D.C., Hu, J., Gao, S.(2019) Nat Commun 10: 4914-4914
- PubMed: 31664033 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-12912-0
- Primary Citation Related Structures:
6JFK, 6JFL, 6JFM - PubMed Abstract:
Mitofusin-2 (MFN2) is a dynamin-like GTPase that plays a central role in regulating mitochondrial fusion and cell metabolism. Mutations in MFN2 cause the neurodegenerative disease Charcot-Marie-Tooth type 2A (CMT2A). The molecular basis underlying the physiological and pathological relevance of MFN2 is unclear. Here, we present crystal structures of truncated human MFN2 in different nucleotide-loading states. Unlike other dynamin superfamily members including MFN1, MFN2 forms sustained dimers even after GTP hydrolysis via the GTPase domain (G) interface, which accounts for its high membrane-tethering efficiency. The biochemical discrepancy between human MFN2 and MFN1 largely derives from a primate-only single amino acid variance. MFN2 and MFN1 can form heterodimers via the G interface in a nucleotide-dependent manner. CMT2A-related mutations, mapping to different functional zones of MFN2, lead to changes in GTP hydrolysis and homo/hetero-association ability. Our study provides fundamental insight into how mitofusins mediate mitochondrial fusion and the ways their disruptions cause disease.
- State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Sun Yat-sen University Cancer Center, 510060, Guangzhou, China.
Organizational Affiliation: