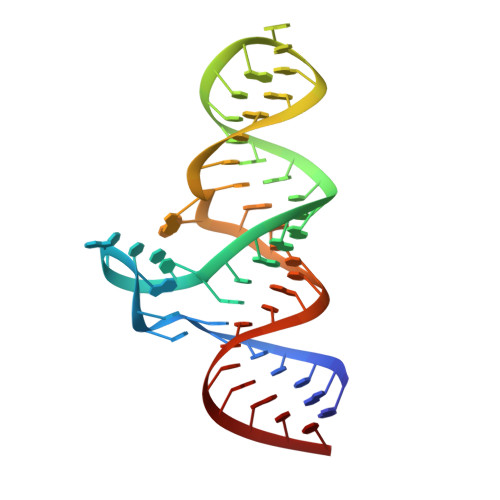
Structure and ligand binding of the ADP-binding domain of the NAD + riboswitch.
Huang, L., Wang, J., Lilley, D.M.J.(2020) RNA 26: 878-887
- PubMed: 32295864
- DOI: https://doi.org/10.1261/rna.074898.120
- Primary Citation Related Structures:
6TB7, 6TF0, 6TF1, 6TF2, 6TF3, 6TFE, 6TFF, 6TFG, 6TFH - PubMed Abstract:
The nad A motif is the first known NAD + -dependent riboswitch, comprising two similar tandem bulged stem-loop structures. We have determined the structure of the 5' domain 1 of the riboswitch. It has three coaxial helical segments, separated by an ACANCCCC bulge and by an internal loop, with a tertiary contact between them that includes two C:G base pairs. We have determined the structure with a number of ligands related to NADH, but in each case only the ADP moiety is observed. The adenosine adopts an anti conformation, forms multiple hydrogen bonds across the width of the sugar edge of the penultimate C:G base pair of the helix preceding the bulge, and the observed contacts have been confirmed by mutagenesis and calorimetry. Two divalent metal ions play a key structural role at the narrow neck of the bulge. One makes direct bonding contacts to the diphosphate moiety, locking it into position. Thus the nucleobase, ribose, and phosphate groups of the ADP moiety are all specifically recognized by the RNA. The NAD + riboswitch is modular. Domain 1 is an ADP binding domain that may be ancient and could potentially be used in combination with other ligand binding motifs such as CoA.
- Cancer Research UK Nucleic Acid Structure Research Group, MSI/WTB Complex, The University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: