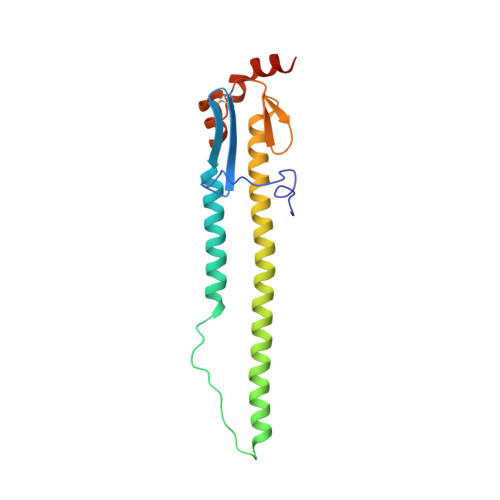
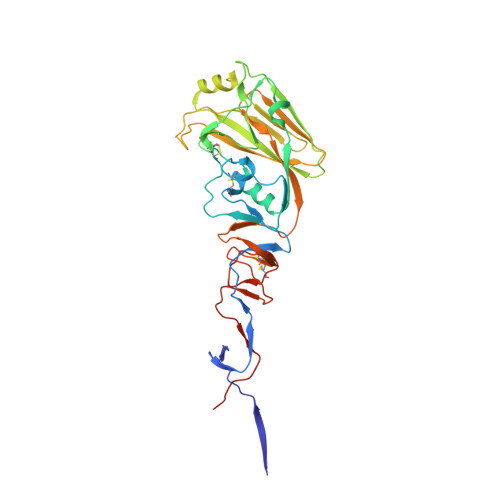An influenza A hemagglutinin small-molecule fusion inhibitor identified by a new high-throughput fluorescence polarization screen.
Yao, Y., Kadam, R.U., Lee, C.D., Woehl, J.L., Wu, N.C., Zhu, X., Kitamura, S., Wilson, I.A., Wolan, D.W.(2020) Proc Natl Acad Sci U S A 117: 18431-18438
- PubMed: 32690700 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2006893117
- Primary Citation Related Structures:
6WCR - PubMed Abstract:
Influenza hemagglutinin (HA) glycoprotein is the primary surface antigen targeted by the host immune response and a focus for development of novel vaccines, broadly neutralizing antibodies (bnAbs), and therapeutics. HA enables viral entry into host cells via receptor binding and membrane fusion and is a validated target for drug discovery. However, to date, only a very few bona fide small molecules have been reported against the HA. To identity new antiviral lead candidates against the highly conserved fusion machinery in the HA stem, we synthesized a fluorescence-polarization probe based on a recently described neutralizing cyclic peptide P7 derived from the complementarity-determining region loops of human bnAbs FI6v3 and CR9114 against the HA stem. We then designed a robust binding assay compatible with high-throughput screening to identify molecules with low micromolar to nanomolar affinity to influenza A group 1 HAs. Our simple, low-cost, and efficient in vitro assay was used to screen H1/Puerto Rico/8/1934 (H1/PR8) HA trimer against ∼72,000 compounds. The crystal structure of H1/PR8 HA in complex with our best hit compound F0045(S) confirmed that it binds to pockets in the HA stem similar to bnAbs FI6v3 and CR9114, cyclic peptide P7, and small-molecule inhibitor JNJ4796. F0045 is enantioselective against a panel of group 1 HAs and F0045(S) exhibits in vitro neutralization activity against multiple H1N1 and H5N1 strains. Our assay, compound characterization, and small-molecule candidate should further stimulate the discovery and development of new compounds with unique chemical scaffolds and enhanced influenza antiviral capabilities.
- Department of Molecular Medicine, The Scripps Research Institute, La Jolla, CA 92037.
Organizational Affiliation: