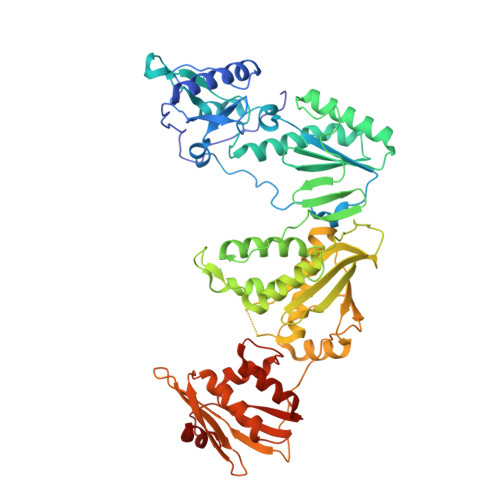
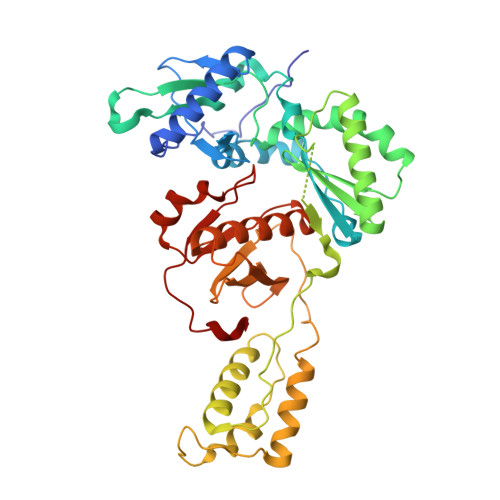
Covalent Inhibition of Wild-Type HIV-1 Reverse Transcriptase Using a Fluorosulfate Warhead.
Ippolito, J.A., Niu, H., Bertoletti, N., Carter, Z.J., Jin, S., Spasov, K.A., Cisneros, J.A., Valhondo, M., Cutrona, K.J., Anderson, K.S., Jorgensen, W.L.(2021) ACS Med Chem Lett 12: 249-255
- PubMed: 33603971 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.0c00612
- Primary Citation Related Structures:
7KRC, 7KRD, 7KRE, 7KRF - PubMed Abstract:
Covalent inhibitors of wild-type HIV-1 reverse transcriptase (CRTIs) are reported. Three compounds derived from catechol diether non-nucleoside inhibitors (NNRTIs) with addition of a fluorosulfate warhead are demonstrated to covalently modify Tyr181 of HIV-RT. X-ray crystal structures for complexes of the CRTIs with the enzyme are provided, which fully demonstrate the covalent attachment, and confirmation is provided by appropriate mass shifts in ESI-TOF mass spectra. The three CRTIs and six noncovalent analogues are found to be potent inhibitors with both IC 50 values for in vitro inhibition of WT RT and EC 50 values for cytopathic protection of HIV-1-infected human T-cells in the 5-320 nM range.
- Department of Chemistry, Yale University, New Haven, Connecticut 06520-8107, United States.
Organizational Affiliation: