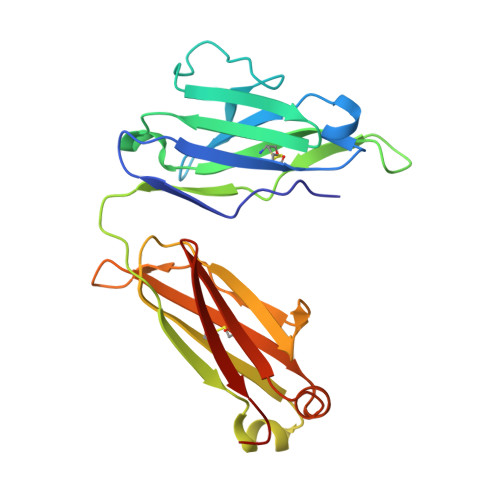
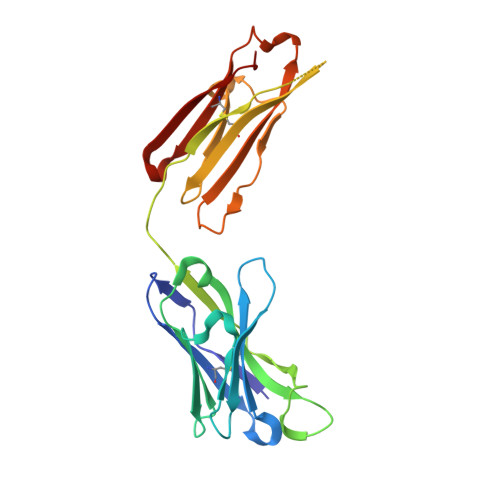
SARS-CoV-2 RBD antibodies that maximize breadth and resistance to escape.
Starr, T.N., Czudnochowski, N., Liu, Z., Zatta, F., Park, Y.J., Addetia, A., Pinto, D., Beltramello, M., Hernandez, P., Greaney, A.J., Marzi, R., Glass, W.G., Zhang, I., Dingens, A.S., Bowen, J.E., Tortorici, M.A., Walls, A.C., Wojcechowskyj, J.A., De Marco, A., Rosen, L.E., Zhou, J., Montiel-Ruiz, M., Kaiser, H., Dillen, J.R., Tucker, H., Bassi, J., Silacci-Fregni, C., Housley, M.P., di Iulio, J., Lombardo, G., Agostini, M., Sprugasci, N., Culap, K., Jaconi, S., Meury, M., Dellota Jr., E., Abdelnabi, R., Foo, S.C., Cameroni, E., Stumpf, S., Croll, T.I., Nix, J.C., Havenar-Daughton, C., Piccoli, L., Benigni, F., Neyts, J., Telenti, A., Lempp, F.A., Pizzuto, M.S., Chodera, J.D., Hebner, C.M., Virgin, H.W., Whelan, S.P.J., Veesler, D., Corti, D., Bloom, J.D., Snell, G.(2021) Nature 597: 97-102
- PubMed: 34261126 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03807-6
- Primary Citation Related Structures:
7M7W, 7R6W, 7R6X, 7R7N - PubMed Abstract:
An ideal therapeutic anti-SARS-CoV-2 antibody would resist viral escape 1-3 , have activity against diverse sarbecoviruses 4-7 , and be highly protective through viral neutralization 8-11 and effector functions 12,13 . Understanding how these properties relate to each other and vary across epitopes would aid the development of therapeutic antibodies and guide vaccine design. Here we comprehensively characterize escape, breadth and potency across a panel of SARS-CoV-2 antibodies targeting the receptor-binding domain (RBD). Despite a trade-off between in vitro neutralization potency and breadth of sarbecovirus binding, we identify neutralizing antibodies with exceptional sarbecovirus breadth and a corresponding resistance to SARS-CoV-2 escape. One of these antibodies, S2H97, binds with high affinity across all sarbecovirus clades to a cryptic epitope and prophylactically protects hamsters from viral challenge. Antibodies that target the angiotensin-converting enzyme 2 (ACE2) receptor-binding motif (RBM) typically have poor breadth and are readily escaped by mutations despite high neutralization potency. Nevertheless, we also characterize a potent RBM antibody (S2E12 8 ) with breadth across sarbecoviruses related to SARS-CoV-2 and a high barrier to viral escape. These data highlight principles underlying variation in escape, breadth and potency among antibodies that target the RBD, and identify epitopes and features to prioritize for therapeutic development against the current and potential future pandemics.
- Basic Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, WA, USA.
Organizational Affiliation: