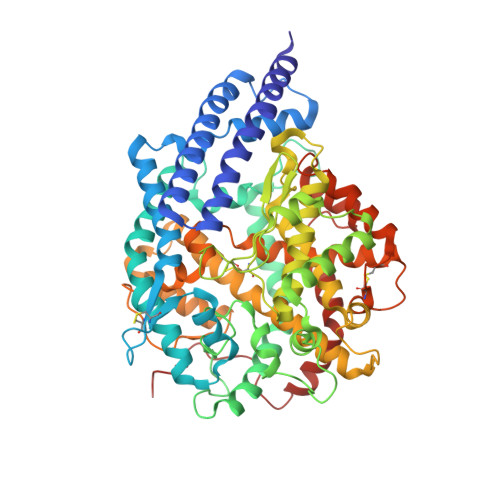
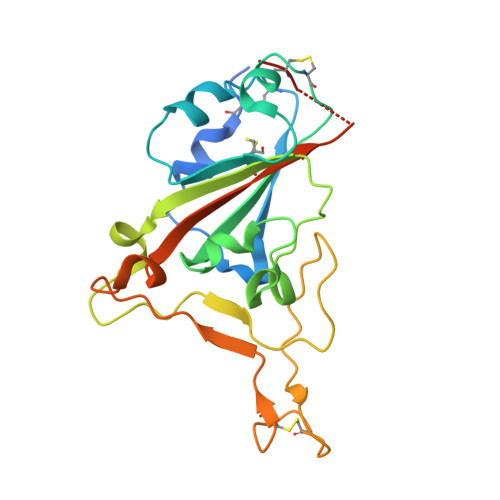
Engineered ACE2-Fc counters murine lethal SARS-CoV-2 infection through direct neutralization and Fc-effector activities.
Chen, Y., Sun, L., Ullah, I., Beaudoin-Bussieres, G., Anand, S.P., Hederman, A.P., Tolbert, W.D., Sherburn, R., Nguyen, D.N., Marchitto, L., Ding, S., Wu, D., Luo, Y., Gottumukkala, S., Moran, S., Kumar, P., Piszczek, G., Mothes, W., Ackerman, M.E., Finzi, A., Uchil, P.D., Gonzalez, F.J., Pazgier, M.(2021) bioRxiv
- PubMed: 34845451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2021.11.24.469776
- Primary Citation Related Structures:
7RPV - PubMed Abstract:
Soluble Angiotensin-Converting Enzyme 2 (ACE2) constitutes an attractive antiviral capable of targeting a wide range of coronaviruses utilizing ACE2 as their receptor. Here, using structure-guided approaches, we developed divalent ACE2 molecules by grafting the extracellular ACE2-domain onto a human IgG1 or IgG3 (ACE2-Fc). These ACE2-Fcs harbor structurally validated mutations that enhance spike (S) binding and remove angiotensin enzymatic activity. The lead variant bound tightly to S, mediated in vitro neutralization of SARS-CoV-2 variants of concern (VOCs) with sub-nanomolar IC 50 and was capable of robust Fc-effector functions, including antibody-dependent-cellular cytotoxicity, phagocytosis and complement deposition. When tested in a stringent K18-hACE2 mouse model, it delayed death or effectively resolved lethal SARS-CoV-2 infection in a prophylactic or therapeutic setting utilizing the combined effect of neutralization and Fc-effector functions. These data confirm the utility of ACE2-Fcs as valuable agents in preventing and eliminating SARS-CoV-2 infection and demonstrate that ACE2-Fc therapeutic activity require Fc-effector functions.
- Infectious Disease Division, Department of Medicine, Uniformed Services University of the Health Sciences, Bethesda, MD 20814-4712, USA.
Organizational Affiliation: