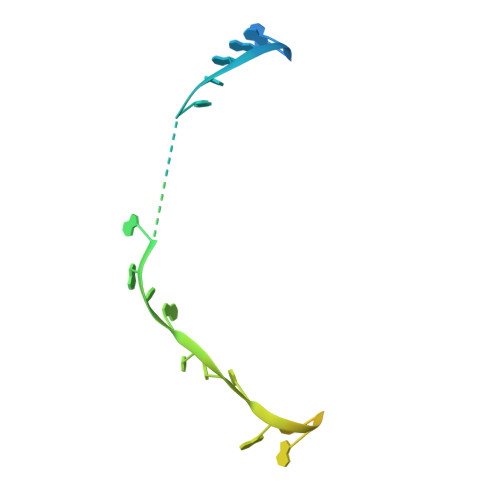
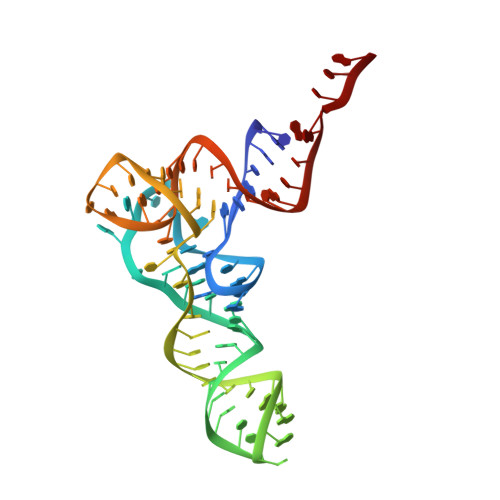
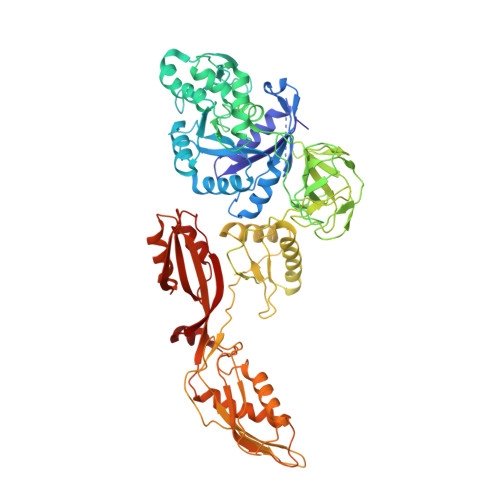
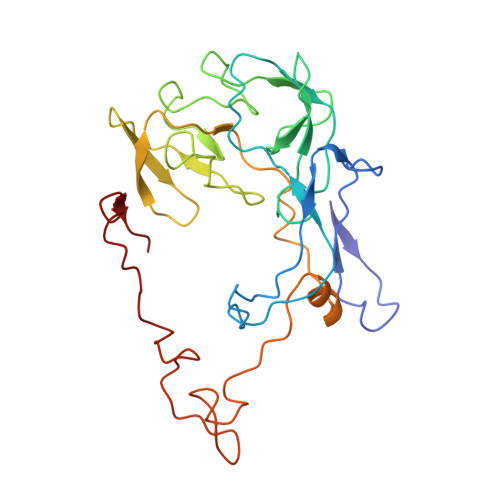
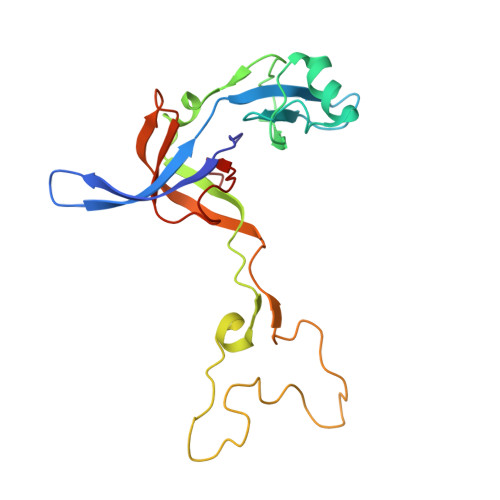
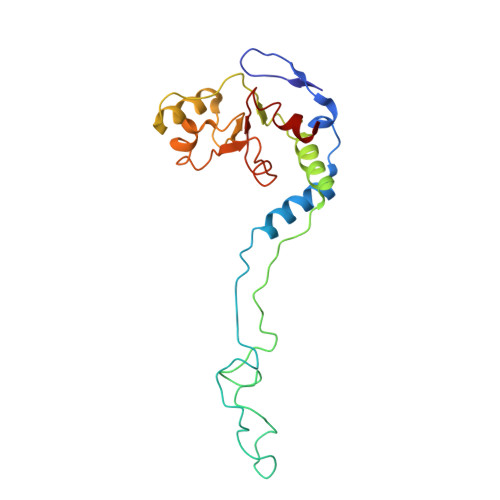
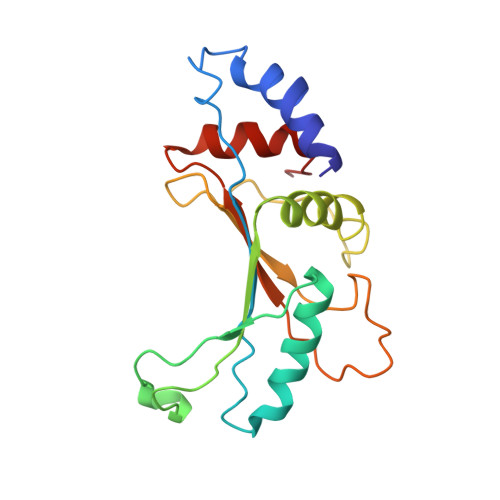
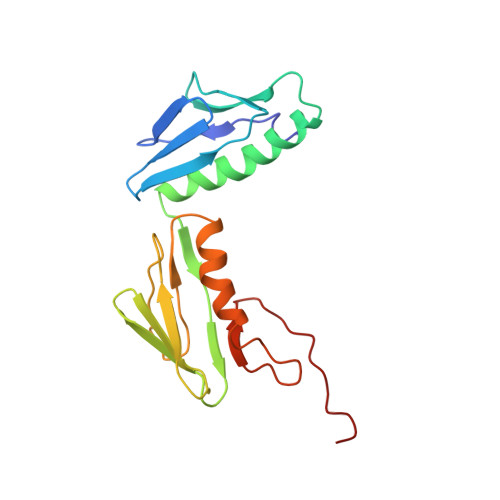
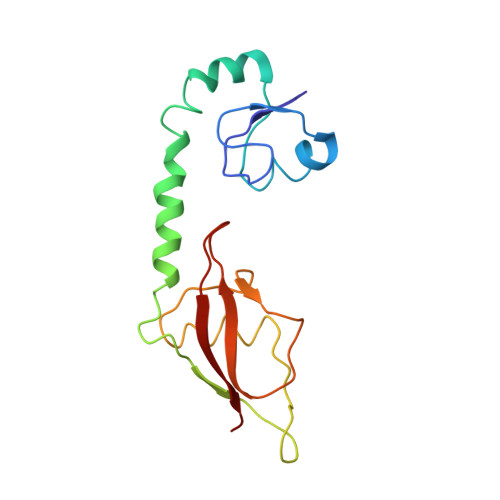
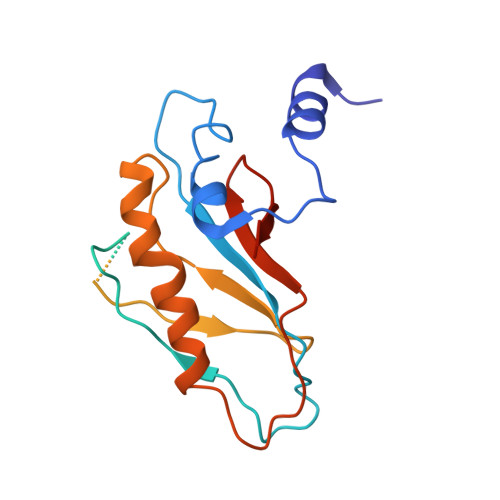
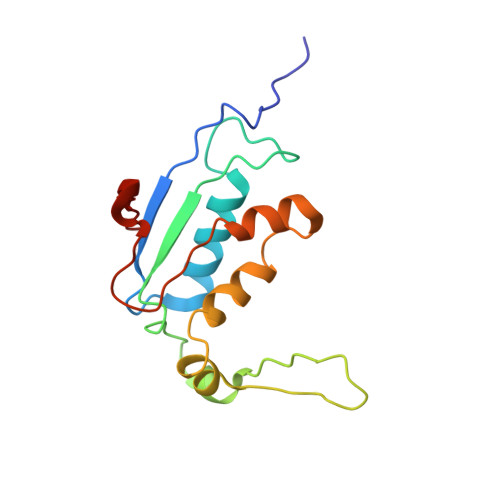
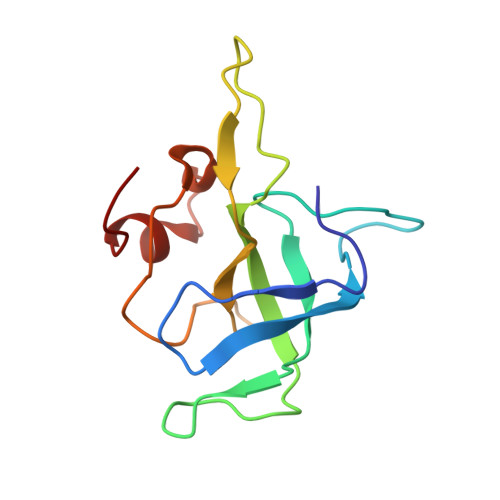
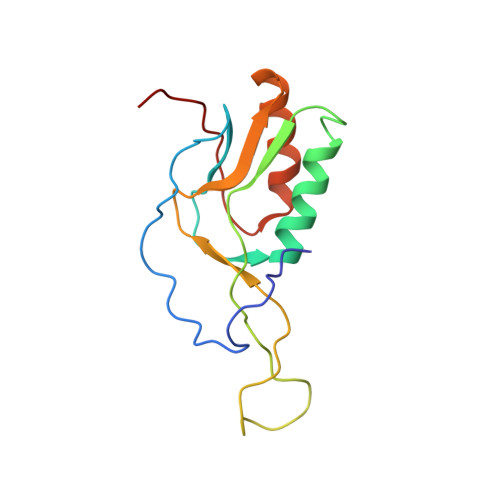
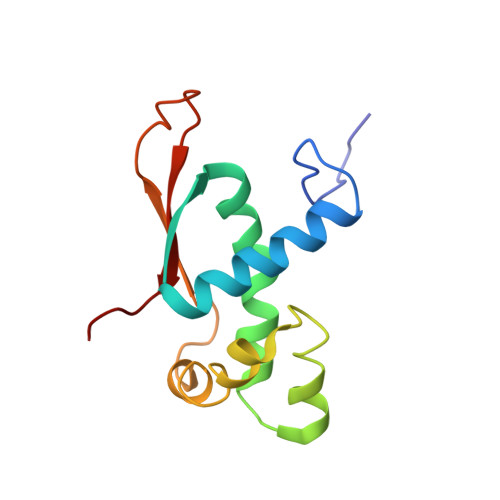
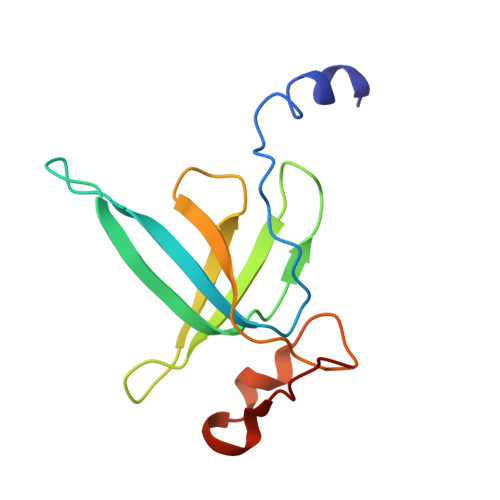
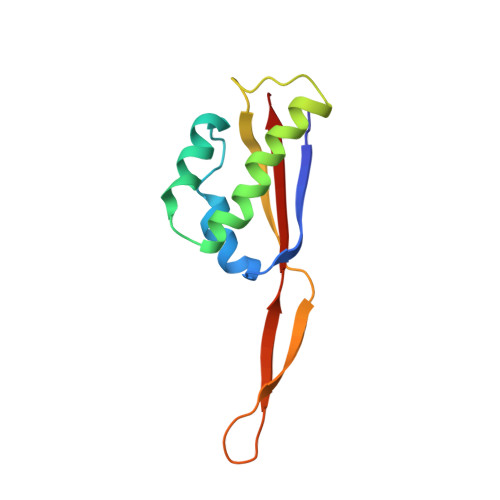
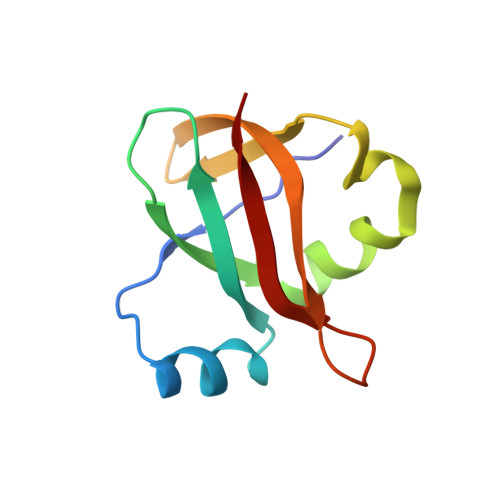
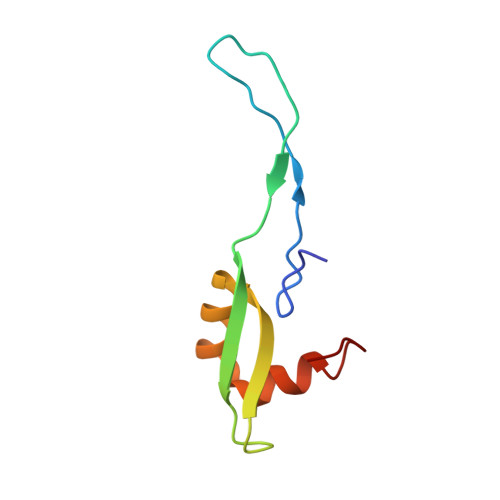
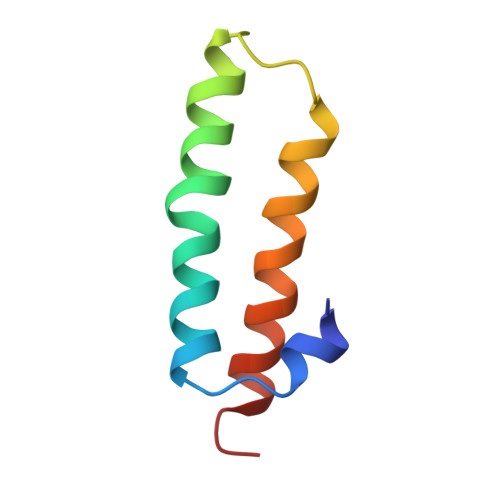
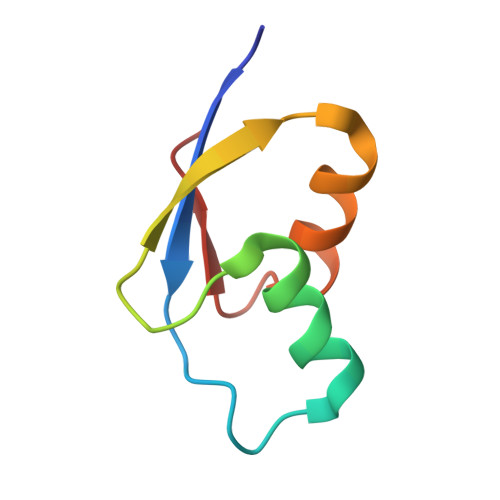
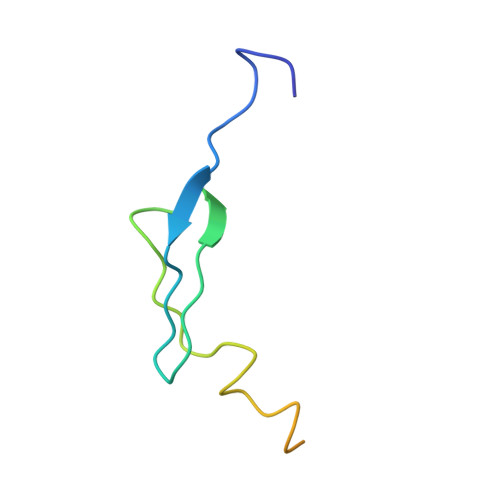
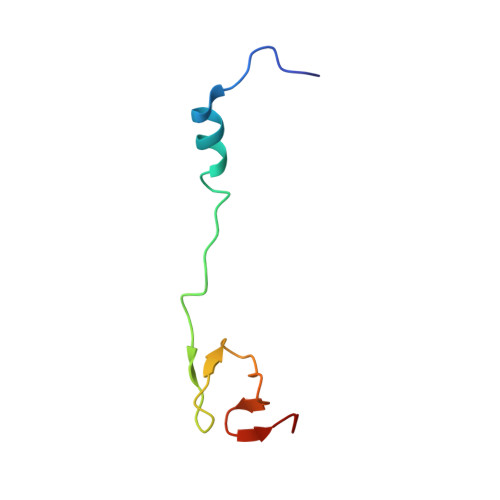
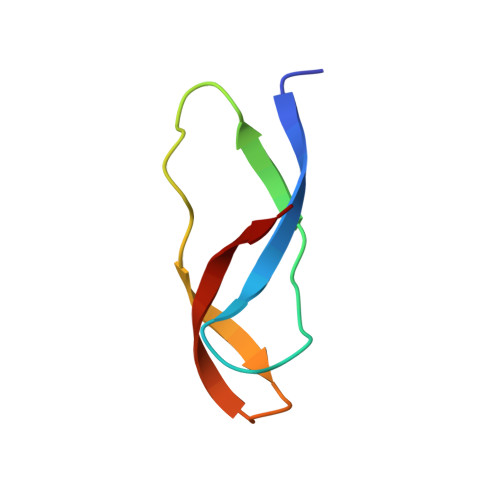
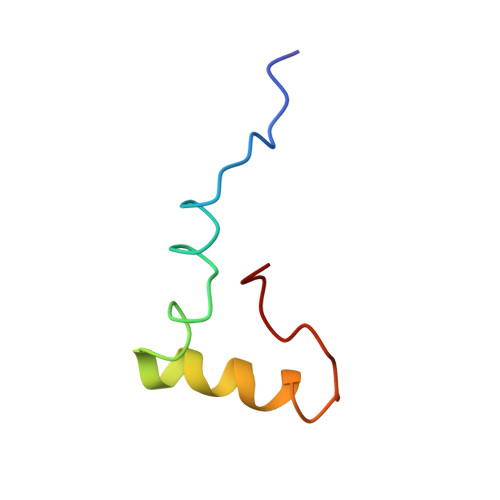
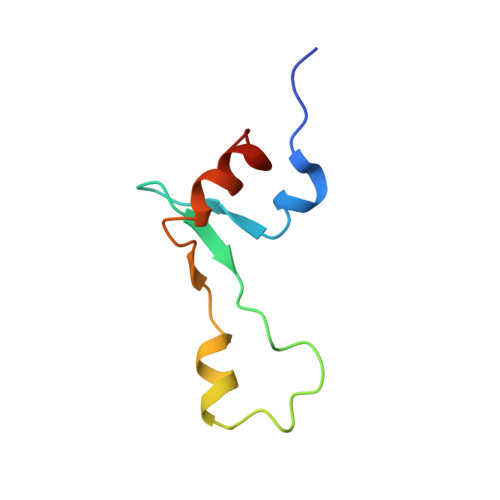
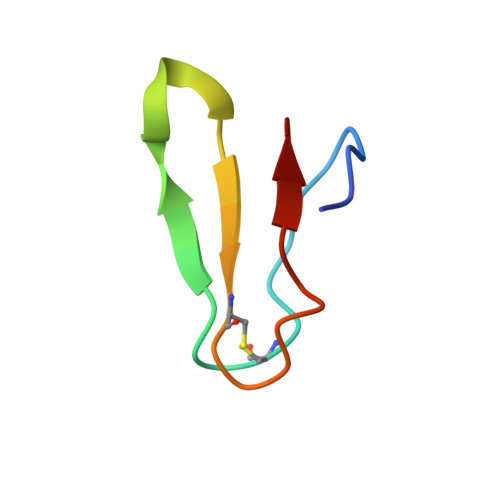
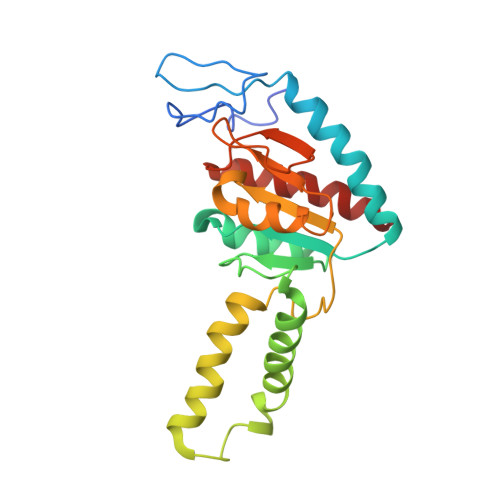
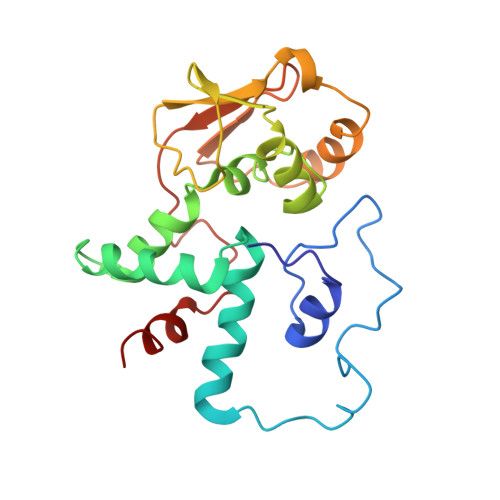
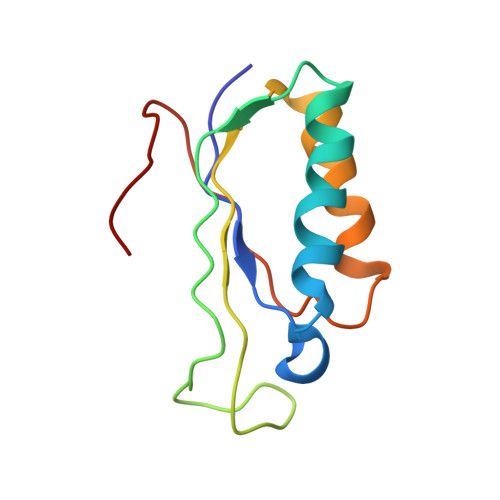
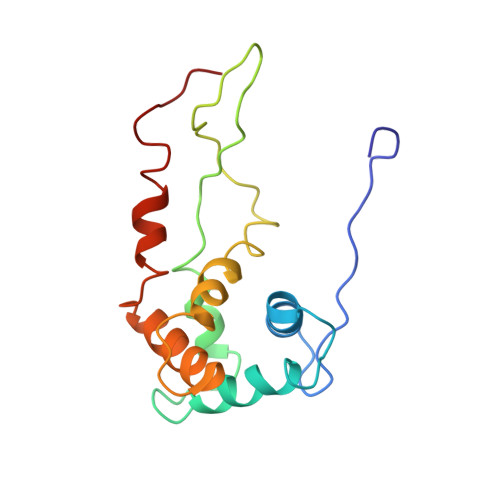
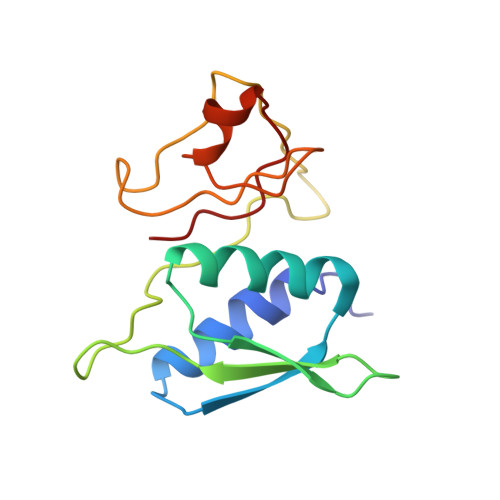
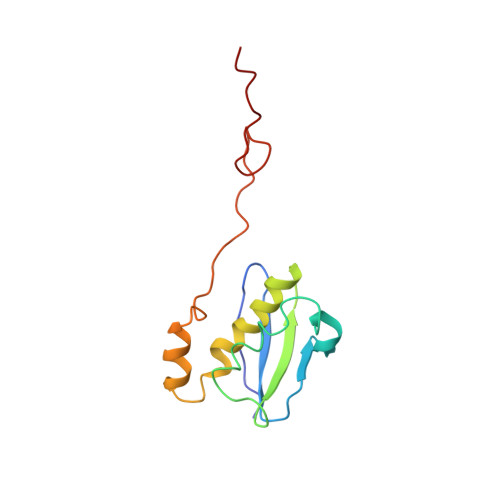
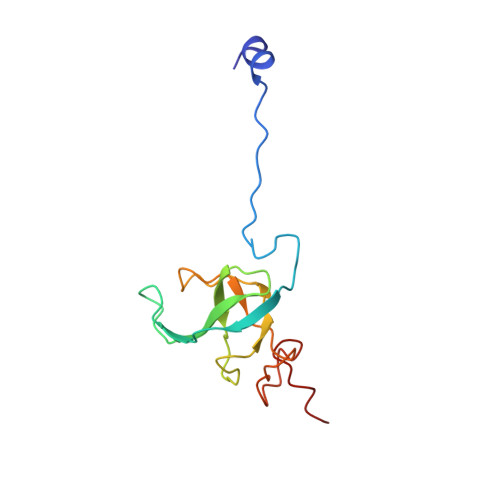
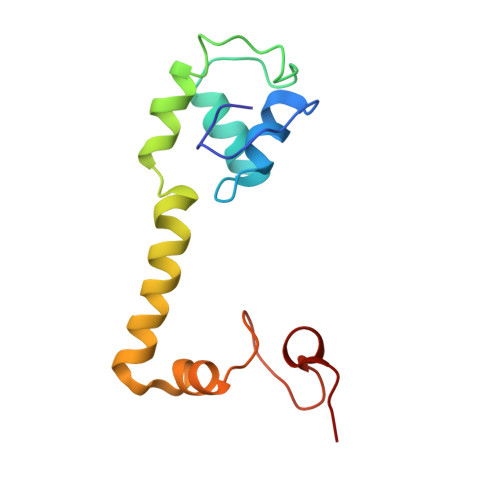
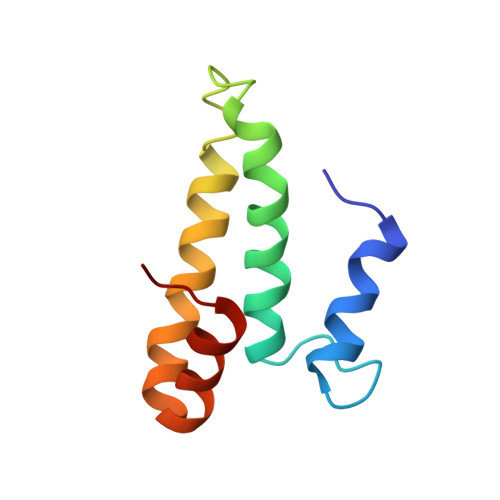
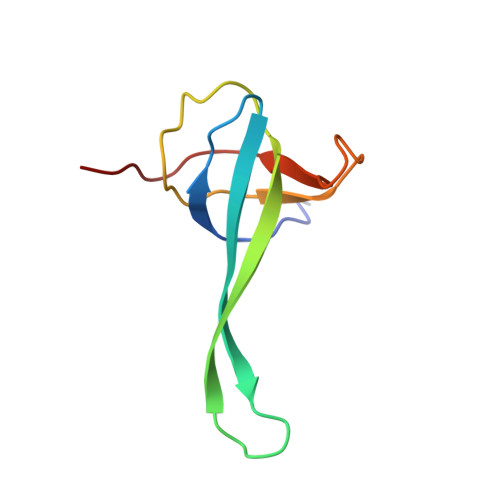
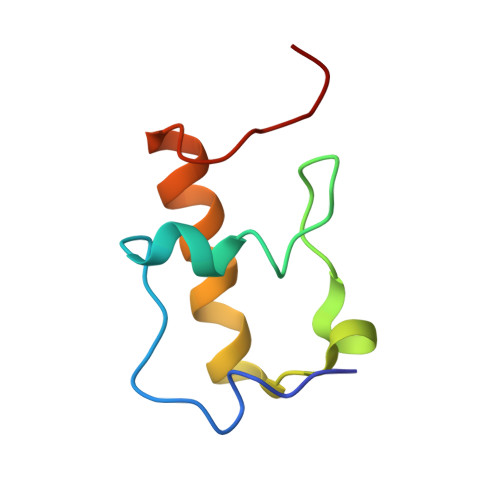
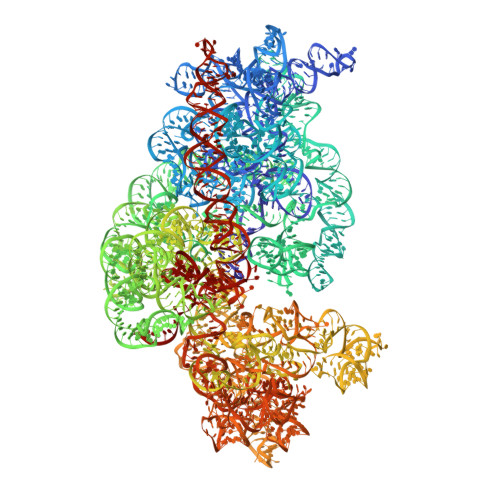
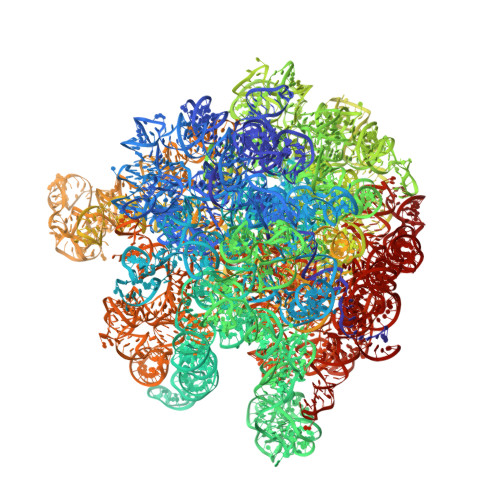
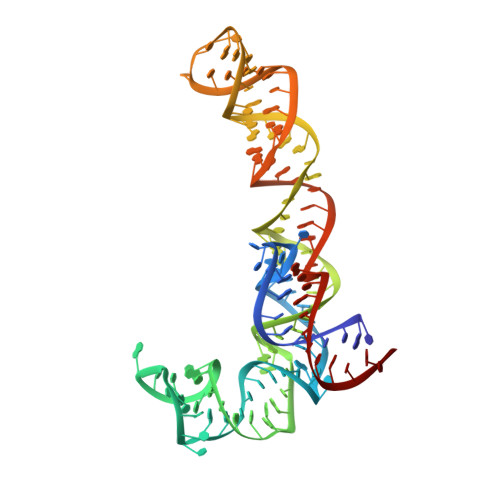
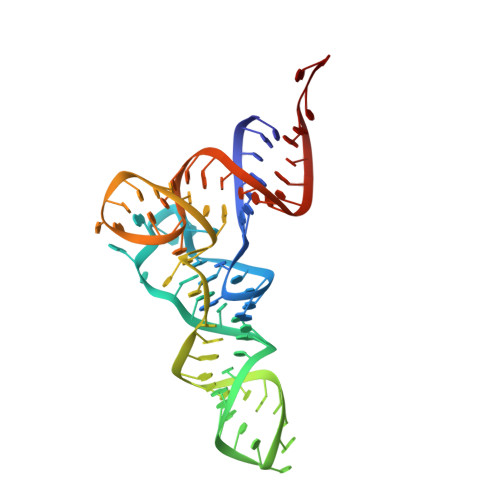Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP.
Carbone, C.E., Loveland, A.B., Gamper Jr., H.B., Hou, Y.M., Demo, G., Korostelev, A.A.(2021) Nat Commun 12: 7236-7236
- PubMed: 34903725 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27415-0
- Primary Citation Related Structures:
7SS9, 7SSD, 7SSL, 7SSN, 7SSO, 7SSW, 7ST2, 7ST6, 7ST7 - PubMed Abstract:
During translation, a conserved GTPase elongation factor-EF-G in bacteria or eEF2 in eukaryotes-translocates tRNA and mRNA through the ribosome. EF-G has been proposed to act as a flexible motor that propels tRNA and mRNA movement, as a rigid pawl that biases unidirectional translocation resulting from ribosome rearrangements, or by various combinations of motor- and pawl-like mechanisms. Using time-resolved cryo-EM, we visualized GTP-catalyzed translocation without inhibitors, capturing elusive structures of ribosome•EF-G intermediates at near-atomic resolution. Prior to translocation, EF-G binds near peptidyl-tRNA, while the rotated 30S subunit stabilizes the EF-G GTPase center. Reverse 30S rotation releases Pi and translocates peptidyl-tRNA and EF-G by ~20 Å. An additional 4-Å translocation initiates EF-G dissociation from a transient ribosome state with highly swiveled 30S head. The structures visualize how nearly rigid EF-G rectifies inherent and spontaneous ribosomal dynamics into tRNA-mRNA translocation, whereas GTP hydrolysis and Pi release drive EF-G dissociation.
- RNA Therapeutics Institute, UMass Chan Medical School, Worcester, MA, USA.
Organizational Affiliation: