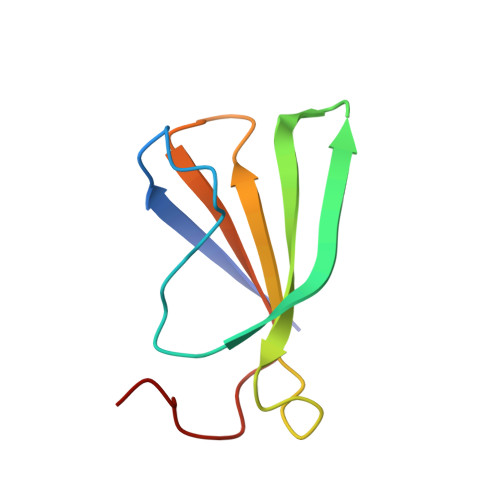
New aspects of DNA recognition by group II WRKY transcription factor revealed by structural and functional study of AtWRKY18 DNA binding domain.
Grzechowiak, M., Ruszkowska, A., Sliwiak, J., Urbanowicz, A., Jaskolski, M., Ruszkowski, M.(2022) Int J Biol Macromol 213: 589-601
- PubMed: 35660042 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2022.05.186
- Primary Citation Related Structures:
7Z0R, 7Z0U - PubMed Abstract:
WRKY transcription factors (TFs) constitute one of the largest families of plant TFs. Based on the organization of domains and motifs, WRKY TFs are divided into three Groups (I-III). The WRKY subgroup IIa includes three representatives in A. thaliana, AtWRKY18, AtWRKY40, and AtWRKY60, that participate in biotic and abiotic stress responses. Here we present crystal structures of the DNA binding domain (DBD) of AtWRKY18 alone and in the complex with a DNA duplex containing the WRKY-recognition sequence, W-box. Subgroup IIa WRKY TFs are known to form homo and heterodimers. Our data suggest that the dimerization interface of the full-length AtWRKY18 involves contacts between the DBD subunits. DNA binding experiments and structural analysis point out novel aspects of DNA recognition by WRKY TFs. In particular, AtWRKY18-DBD preferentially binds an overlapping tandem of W-boxes accompanied by a quasi-W-box motif. The binding of DNA deforms the B-type double helix, which suggests that the DNA fragment must be prone to form a specific structure. This can explain why despite the short W-box consensus, WRKY TFs can precisely control gene expression. Finally, this first experimental structure of a Group II WRKY TF allowed us to compare Group I-III representatives.
- Department of Structural Biology of Eukaryotes, Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan 61-704, Poland.
Organizational Affiliation: