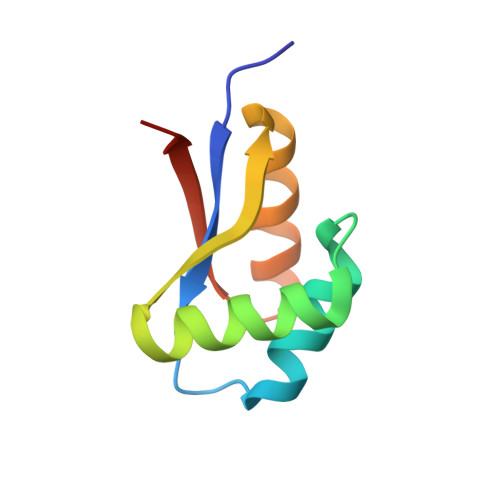
Structural basis for the N-degron specificity of ClpS1 from Arabidopsis thaliana.
Kim, L., Heo, J., Kwon, D.H., Shin, J.S., Jang, S.H., Park, Z.Y., Song, H.K.(2021) Protein Sci 30: 700-708
- PubMed: 33368743 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4018
- Primary Citation Related Structures:
7D34 - PubMed Abstract:
The N-degron pathway determines the half-life of proteins in both prokaryotes and eukaryotes by precisely recognizing the N-terminal residue (N-degron) of substrates. ClpS proteins from bacteria bind to substrates containing hydrophobic N-degrons (Leu, Phe, Tyr, and Trp) and deliver them to the caseinolytic protease system ClpAP. This mechanism is preserved in organelles such as mitochondria and chloroplasts. Bacterial ClpS adaptors bind preferentially to Leu and Phe N-degrons; however, ClpS1 from Arabidopsis thaliana (AtClpS1) shows a difference in that it binds strongly to Phe and Trp N-degrons and only weakly to Leu. This difference in behavior cannot be explained without structural information due to the high sequence homology between bacterial and plant ClpS proteins. Here, we report the structure of AtClpS1 at 2.0 Å resolution in the presence of a bound N-degron. The key determinants for α-amino group recognition are conserved among all ClpS proteins, but the α3-helix of eukaryotic AtClpS1 is significantly shortened, and consequently, a loop forming a pocket for the N-degron is moved slightly outward to enlarge the pocket. In addition, amino acid replacement from Val to Ala causes a reduction in hydrophobic interactions with Leu N-degron. A combination of the fine-tuned hydrophobic residues in the pocket and the basic gatekeeper at the entrance of the pocket controls the N-degron selectivity of the plant ClpS protein.
- Department of Life Sciences, Korea University, Seoul, South Korea.
Organizational Affiliation: